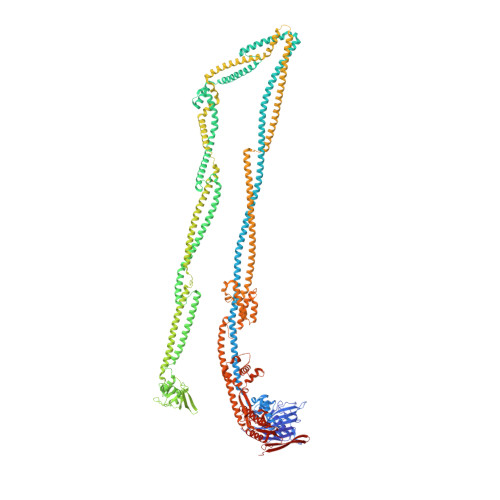
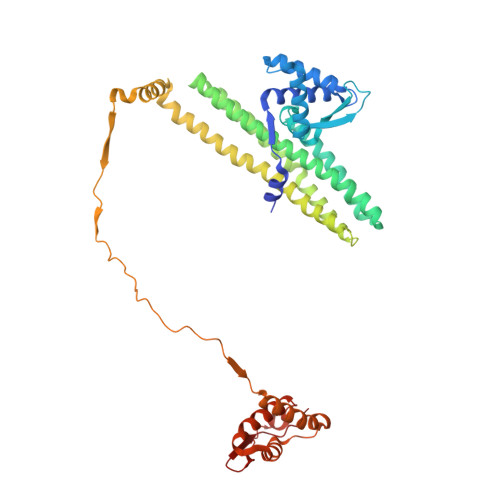
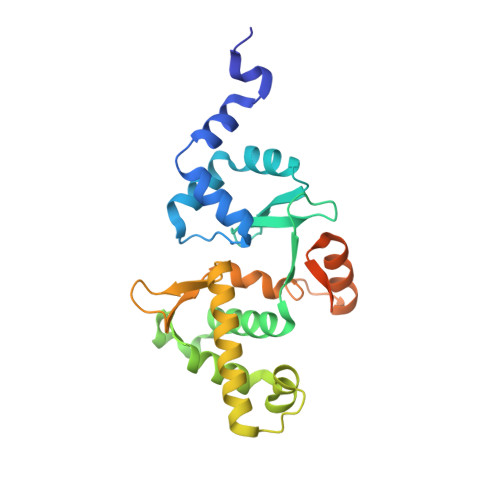
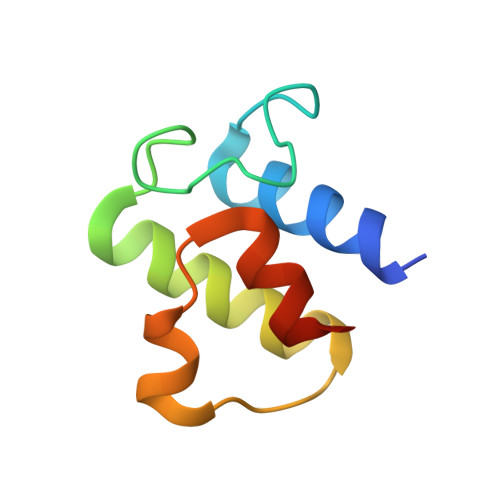
Cryo-EM structure of MukBEF reveals DNA loop entrapment at chromosomal unloading sites.
Burmann, F., Funke, L.F.H., Chin, J.W., Lowe, J.(2021) Mol Cell 81: 4891-4906.e8
- PubMed: 34739874
- DOI: https://doi.org/10.1016/j.molcel.2021.10.011
- Primary Citation of Related Structures:
7NYW, 7NYX, 7NYY, 7NYZ, 7NZ0, 7NZ2, 7NZ3, 7NZ4 - PubMed Abstract:
The ring-like structural maintenance of chromosomes (SMC) complex MukBEF folds the genome of Escherichia coli and related bacteria into large loops, presumably by active DNA loop extrusion. MukBEF activity within the replication terminus macrodomain is suppressed by the sequence-specific unloader MatP. Here, we present the complete atomic structure of MukBEF in complex with MatP and DNA as determined by electron cryomicroscopy (cryo-EM). The complex binds two distinct DNA double helices corresponding to the arms of a plectonemic loop. MatP-bound DNA threads through the MukBEF ring, while the second DNA is clamped by the kleisin MukF, MukE, and the MukB ATPase heads. Combinatorial cysteine cross-linking confirms this topology of DNA loop entrapment in vivo. Our findings illuminate how a class of near-ubiquitous DNA organizers with important roles in genome maintenance interacts with the bacterial chromosome.
Organizational Affiliation:
MRC Laboratory of Molecular Biology, Structural Studies Division, Cambridge Biomedical Campus, Cambridge, UK. Electronic address: fburmann@mrc-lmb.cam.ac.uk.