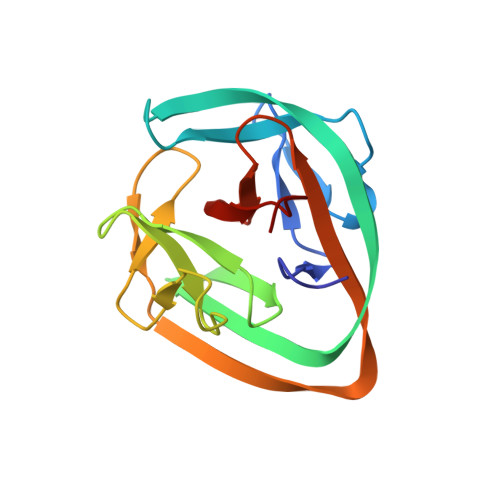
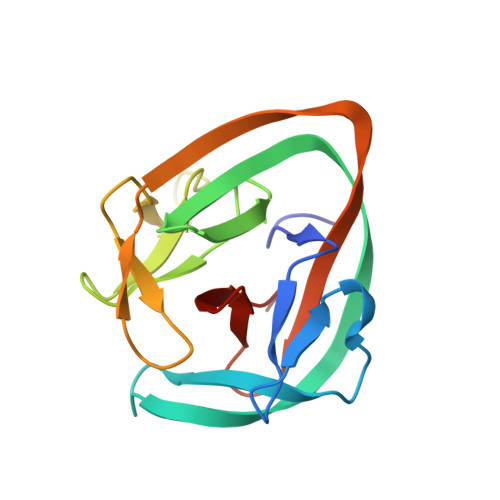
Structural basis of ubiquitination mediated by protein splicing in early Eukarya.
Chiarini, V., Fiorillo, A., Camerini, S., Crescenzi, M., Nakamura, S., Battista, T., Guidoni, L., Colotti, G., Ilari, A.(2021) Biochim Biophys Acta Gen Subj 1865: 129844-129844
- PubMed: 33444728
- DOI: https://doi.org/10.1016/j.bbagen.2021.129844
- Primary Citation of Related Structures:
6TMM, 6Y75 - PubMed Abstract:
Inteins are intervening proteins, which are known to perform protein splicing. The reaction results in the production of an intein domain and an inteinless protein, which shows no trace of the insertion. BIL2 is part of the polyubiquitin locus of Tetrahymena thermophila (BUBL), where two bacterial-intein-like (BIL) domains lacking the C + 1 nucleophile, are flanked by two independent ubiquitin-like domains (ubl4/ubl5).
Organizational Affiliation:
Program in Structural Biology and Biophysics, Institute of Biotechnology, University of Helsinki, Viikinkaari 1, P.O. Box 65, FI-00014 Helsinki, Finland.