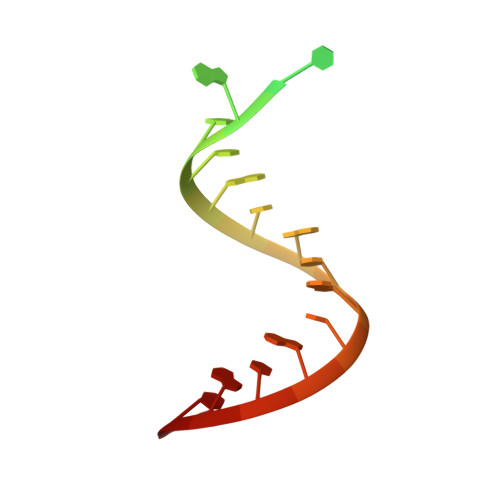
Evolution of Inosine-Specific Endonuclease V from Bacterial DNase to Eukaryotic RNase.
Wu, J., Samara, N.L., Kuraoka, I., Yang, W.(2019) Mol Cell 76: 44
- PubMed: 31444105
- DOI: https://doi.org/10.1016/j.molcel.2019.06.046
- Primary Citation of Related Structures:
6OZE, 6OZF, 6OZG, 6OZH, 6OZI, 6OZJ, 6OZK, 6OZL, 6OZM, 6OZN, 6OZO, 6OZP, 6OZQ, 6OZR, 6OZS - PubMed Abstract:
Endonuclease V (EndoV) cleaves the second phosphodiester bond 3' to a deaminated adenosine (inosine). Although highly conserved, EndoV homologs change substrate preference from DNA in bacteria to RNA in eukaryotes. We have characterized EndoV from six different species and determined crystal structures of human EndoV and three EndoV homologs from bacteria to mouse in complex with inosine-containing DNA/RNA hybrid or double-stranded RNA (dsRNA). Inosine recognition is conserved, but changes in several connecting loops in eukaryotic EndoV confer recognition of 3 ribonucleotides upstream and 7 or 8 bp of dsRNA downstream of the cleavage site, and bacterial EndoV binds only 2 or 3 nt flanking the scissile phosphate. In addition to the two canonical metal ions in the active site, a third Mn 2+ that coordinates the nucleophilic water appears necessary for product formation. Comparison of EndoV with its homologs RNase H1 and Argonaute reveals the principles by which these enzymes recognize RNA versus DNA.
Organizational Affiliation:
Laboratory of Molecular Biology, NIDDK, NIH, Bethesda, MD 20892, USA.