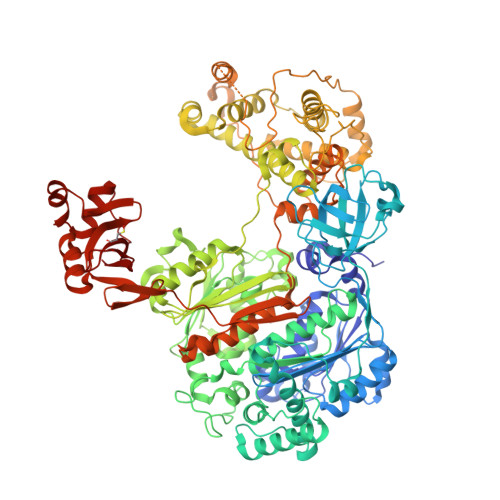
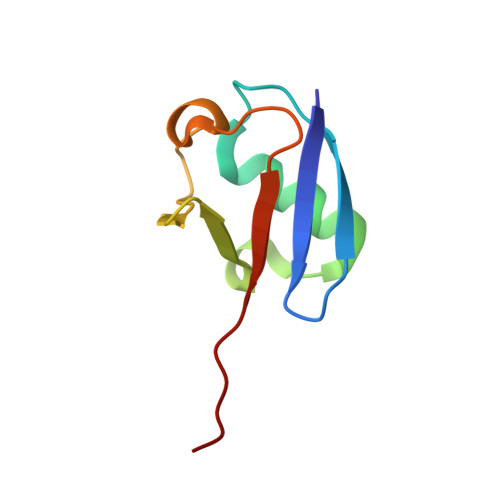
Dissecting the Specificity of Adenosyl Sulfamate Inhibitors Targeting the Ubiquitin-Activating Enzyme.
Misra, M., Kuhn, M., Lobel, M., An, H., Statsyuk, A.V., Sotriffer, C., Schindelin, H.(2017) Structure 25: 1120-1129.e3
- PubMed: 28578874
- DOI: https://doi.org/10.1016/j.str.2017.05.001
- Primary Citation of Related Structures:
5L6H, 5L6I, 5L6J - PubMed Abstract:
Targeting the activating enzymes (E1) of ubiquitin (Ub) and ubiquitin-like modifiers (Ubls) has emerged as a promising anti-cancer strategy, possibly overcoming the ineffectiveness of proteasome inhibitors against solid tumors. Here, we report crystal structures of the yeast ubiquitin E1 (Uba1) with three adenosyl sulfamate inhibitors exhibiting different E1 specificities, which are all covalently linked to ubiquitin. The structures illustrate how the chemically diverse inhibitors are accommodated within the adenylation active site. When compared with the previously reported structures of various E1 enzymes, our structures provide the basis of the preferences of these inhibitors for different Ub/Ubl-activating enzymes. In vitro inhibition assays and molecular dynamics simulations validated the specificities of the inhibitors as deduced from the structures. Taken together, the structures establish a framework for the development of additional compounds targeting E1 enzymes, which will display higher potency and selectivity.
Organizational Affiliation:
Institute of Structural Biology, Rudolf Virchow Center for Experimental Biomedicine, University of Würzburg, Josef-Schneider-Straße 2, 97080 Würzburg, Germany; Institute of Pharmacy and Food Chemistry, Department of Chemistry and Pharmacy, University of Würzburg, Am Hubland, 97074 Würzburg, Germany.