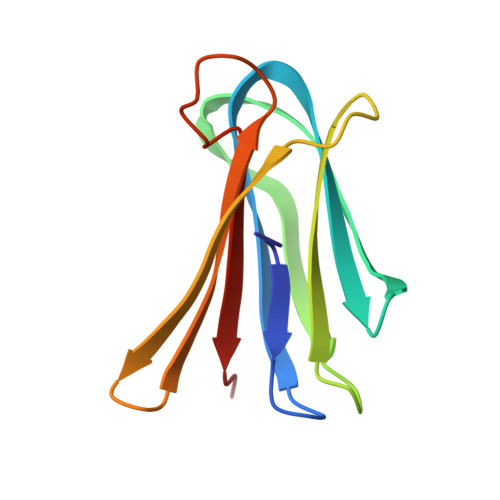
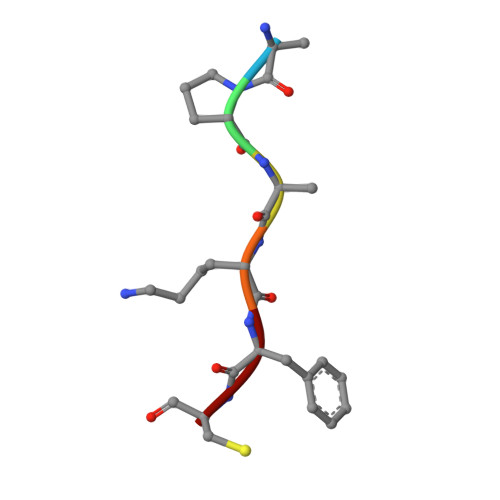
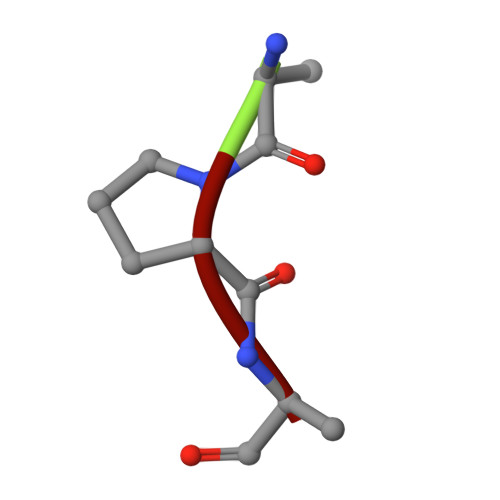
Overcoming antibiotic resistance inPseudomonas aeruginosabiofilms using glycopeptide dendrimers.
Michaud, G., Visini, R., Bergmann, M., Salerno, G., Bosco, R., Gillon, E., Richichi, B., Nativi, C., Imberty, A., Stocker, A., Darbre, T., Reymond, J.L.(2016) Chem Sci 7: 166-182
- PubMed: 29896342
- DOI: https://doi.org/10.1039/c5sc03635f
- Primary Citation of Related Structures:
5D2A - PubMed Abstract:
Antibiotic resistance in the opportunistic pathogen Pseudomonas aeruginosa is partly caused by biofilms forming a physical barrier to antibiotic penetration. Here we focused on modifying tetravalent glycopeptide dendrimer ligands of P. aeruginosa lectins LecB or LecA to increase their biofilm inhibition activity. First heteroglycoclusters were investigated displaying one pair each of LecB specific fucosyl groups and LecA specific galactosyl groups and binding simultaneously to both lectins, one of which gave the first fully resolved crystal structure of a peptide dendrimer as LecB complex providing a structural model for dendrimer-lectin interactions (PDB ; 5D2A). Biofilm inhibition was increased by introducing additional cationic residues in these dendrimers but resulted in bactericidal effects similar to those of non-glycosylated polycationic antimicrobial peptide dendrimers. In a second approach dendrimers displaying four copies of the natural LecB ligand Lewis a were prepared leading to slightly stronger LecB binding and biofilm inhibition. Finally synergistic application of a LecB specific non-bactericidal antibiofilm dendrimer with the antibiotic tobramycin at sub-inhibitory concentrations of both compounds allowed effective biofilm inhibition and dispersal.
Organizational Affiliation:
Department of Chemistry and Biochemistry , University of Berne , Freiestrasse 3 , 3012 Berne , Switzerland . Email: jean-louis.reymond@dcb.unibe.ch.