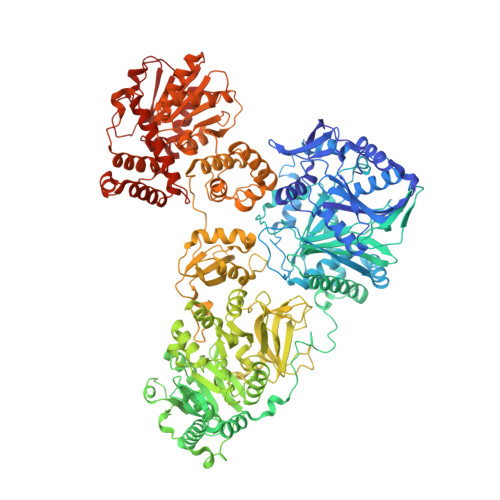
Structures of two distinct conformations of holo-non-ribosomal peptide synthetases.
Drake, E.J., Miller, B.R., Shi, C., Tarrasch, J.T., Sundlov, J.A., Allen, C.L., Skiniotis, G., Aldrich, C.C., Gulick, A.M.(2016) Nature 529: 235-238
- PubMed: 26762461
- DOI: https://doi.org/10.1038/nature16163
- Primary Citation of Related Structures:
4ZXH, 4ZXI, 5T3D - PubMed Abstract:
Many important natural products are produced by multidomain non-ribosomal peptide synthetases (NRPSs). During synthesis, intermediates are covalently bound to integrated carrier domains and transported to neighbouring catalytic domains in an assembly line fashion. Understanding the structural basis for catalysis with non-ribosomal peptide synthetases will facilitate bioengineering to create novel products. Here we describe the structures of two different holo-non-ribosomal peptide synthetase modules, each revealing a distinct step in the catalytic cycle. One structure depicts the carrier domain cofactor bound to the peptide bond-forming condensation domain, whereas a second structure captures the installation of the amino acid onto the cofactor within the adenylation domain. These structures demonstrate that a conformational change within the adenylation domain guides transfer of intermediates between domains. Furthermore, one structure shows that the condensation and adenylation domains simultaneously adopt their catalytic conformations, increasing the overall efficiency in a revised structural cycle. These structures and the single-particle electron microscopy analysis demonstrate a highly dynamic domain architecture and provide the foundation for understanding the structural mechanisms that could enable engineering of novel non-ribosomal peptide synthetases.
Organizational Affiliation:
Hauptman-Woodward Medical Research Institute, 700 Ellicott Street, Buffalo, New York 14203, USA.