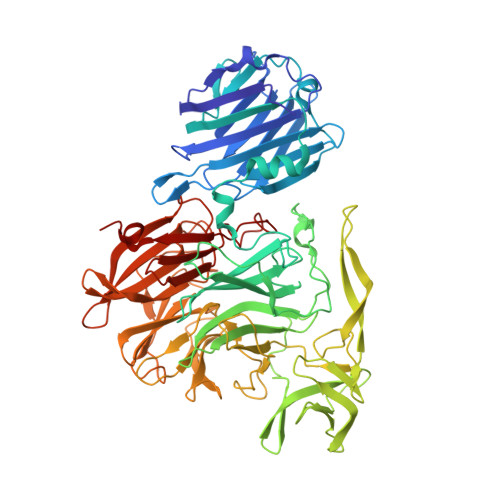
Streptococcus pneumoniae NanC: STRUCTURAL INSIGHTS INTO THE SPECIFICITY AND MECHANISM OF A SIALIDASE THAT PRODUCES A SIALIDASE INHIBITOR.
Owen, C.D., Lukacik, P., Potter, J.A., Sleator, O., Taylor, G.L., Walsh, M.A.(2015) J Biol Chem 290: 27736-27748
- PubMed: 26370075
- DOI: https://doi.org/10.1074/jbc.M115.673632
- Primary Citation of Related Structures:
4YW1, 4YW2, 4YW3, 4YW5, 4YZ1, 4YZ2, 4YZ4, 4YZ5, 5F9T - PubMed Abstract:
Streptococcus pneumoniae is an important human pathogen that causes a range of disease states. Sialidases are important bacterial virulence factors. There are three pneumococcal sialidases: NanA, NanB, and NanC. NanC is an unusual sialidase in that its primary reaction product is 2-deoxy-2,3-didehydro-N-acetylneuraminic acid (Neu5Ac2en, also known as DANA), a nonspecific hydrolytic sialidase inhibitor. The production of Neu5Ac2en from α2-3-linked sialosides by the catalytic domain is confirmed within a crystal structure. A covalent complex with 3-fluoro-β-N-acetylneuraminic acid is also presented, suggesting a common mechanism with other sialidases up to the final step of product formation. A conformation change in an active site hydrophobic loop on ligand binding constricts the entrance to the active site. In addition, the distance between the catalytic acid/base (Asp-315) and the ligand anomeric carbon is unusually short. These features facilitate a novel sialidase reaction in which the final step of product formation is direct abstraction of the C3 proton by the active site aspartic acid, forming Neu5Ac2en. NanC also possesses a carbohydrate-binding module, which is shown to bind α2-3- and α2-6-linked sialosides, as well as N-acetylneuraminic acid, which is captured in the crystal structure following hydration of Neu5Ac2en by NanC. Overall, the pneumococcal sialidases show remarkable mechanistic diversity while maintaining a common structural scaffold.
Organizational Affiliation:
From the Biomedical Sciences Research Complex, University of St. Andrews, St. Andrews, Fife KY16 9ST, United Kingdom.