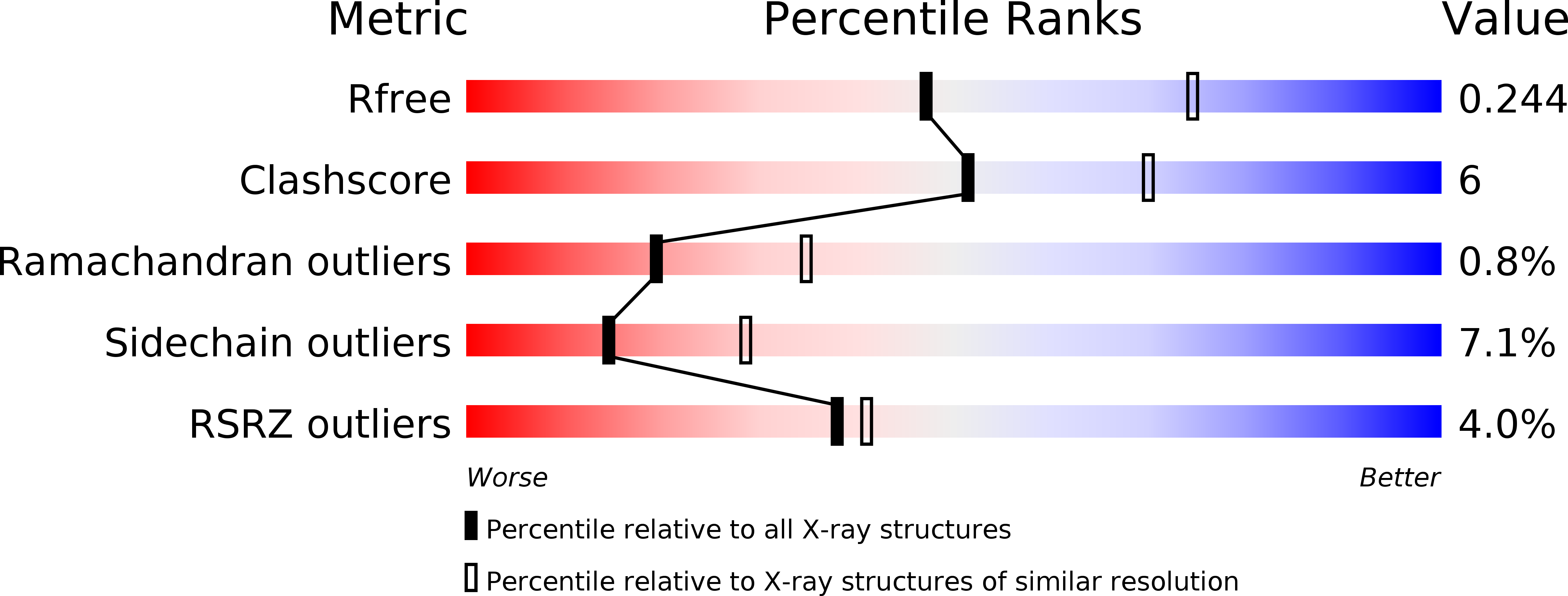
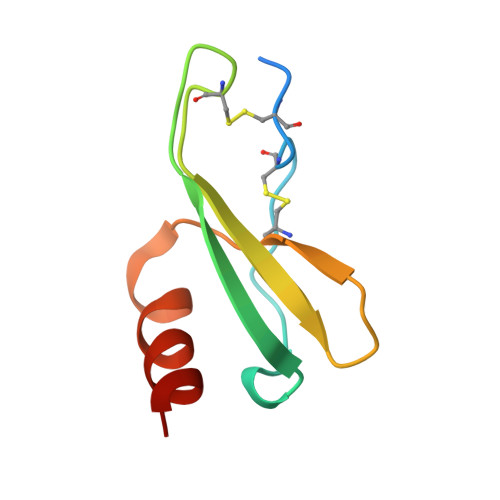
Atomic description of the immune complex involved in heparin-induced thrombocytopenia.
Cai, Z., Yarovoi, S.V., Zhu, Z., Rauova, L., Hayes, V., Lebedeva, T., Liu, Q., Poncz, M., Arepally, G., Cines, D.B., Greene, M.I.(2015) Nat Commun 6: 8277-8277
- PubMed: 26391892
- DOI: https://doi.org/10.1038/ncomms9277
- Primary Citation of Related Structures:
4R97, 4R9W, 4R9Y - PubMed Abstract:
Heparin-induced thrombocytopenia (HIT) is an autoimmune thrombotic disorder caused by immune complexes containing platelet factor 4 (PF4), antibodies to PF4 and heparin or cellular glycosaminoglycans (GAGs). Here we solve the crystal structures of the: (1) PF4 tetramer/fondaparinux complex, (2) PF4 tetramer/KKO-Fab complex (a murine monoclonal HIT-like antibody) and (3) PF4 monomer/RTO-Fab complex (a non-HIT anti-PF4 monoclonal antibody). Fondaparinux binds to the 'closed' end of the PF4 tetramer and stabilizes its conformation. This interaction in turn stabilizes the epitope for KKO on the 'open' end of the tetramer. Fondaparinux and KKO thereby collaborate to 'stabilize' the ternary pathogenic immune complex. Binding of RTO to PF4 monomers prevents PF4 tetramerization and inhibits KKO and human HIT IgG-induced platelet activation and platelet aggregation in vitro, and thrombus progression in vivo. The atomic structures provide a basis to develop new diagnostics and non-anticoagulant therapeutics for HIT.
Organizational Affiliation:
Department of Pathology and Laboratory Medicine, University of Pennsylvania, Philadelphia, Pennsylvania 19104, USA.