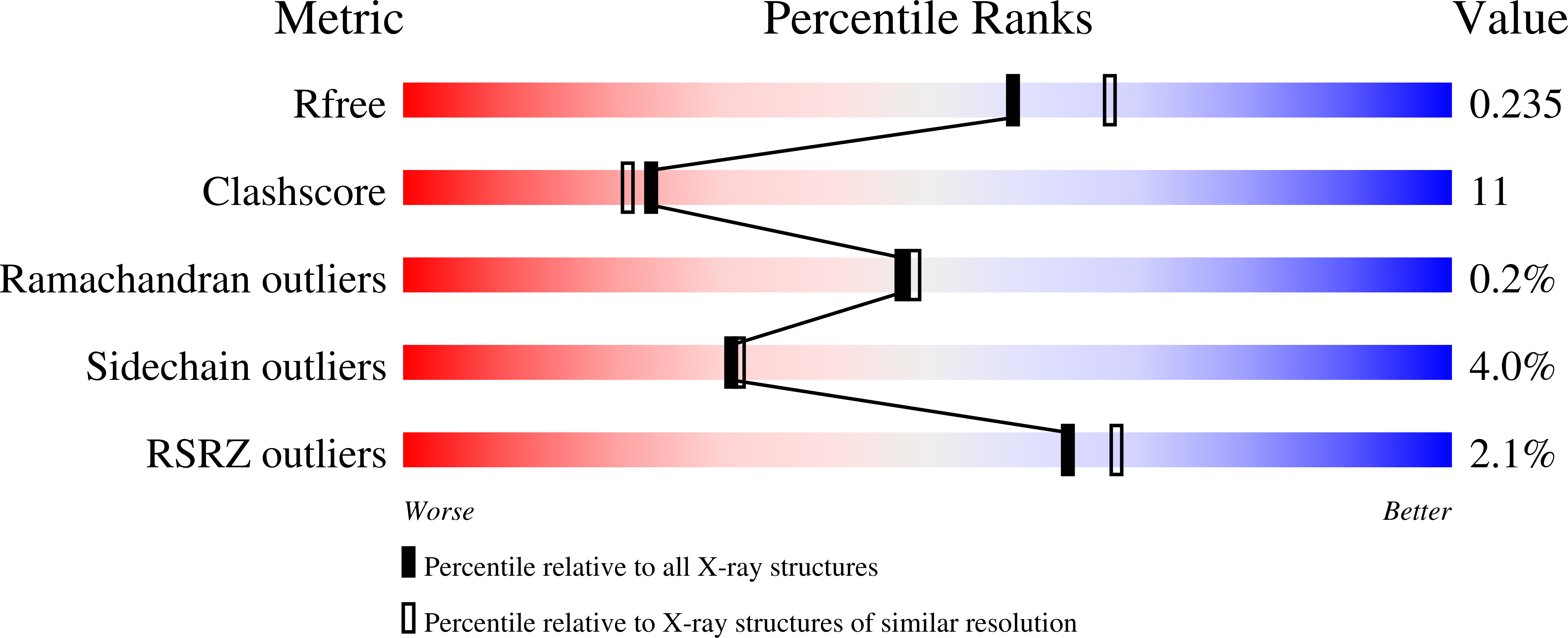
On the catalytic mechanism and stereospecificity of Escherichia coli l-threonine aldolase.
di Salvo, M.L., Remesh, S.G., Vivoli, M., Ghatge, M.S., Paiardini, A., D'Aguanno, S., Safo, M.K., Contestabile, R.(2014) FEBS J 281: 129-145
- PubMed: 24165453
- DOI: https://doi.org/10.1111/febs.12581
- Primary Citation of Related Structures:
4LNJ, 4LNL, 4LNM - PubMed Abstract:
L-threonine aldolases (L-TAs) represent a family of homologous pyridoxal 5'-phosphate-dependent enzymes found in bacteria and fungi, and catalyse the reversible cleavage of several L-3-hydroxy-α-amino acids. L-TAs have great biotechnological potential, as they catalyse the formation of carbon-carbon bonds, and therefore may be exploited for the bioorganic synthesis of L-3-hydroxyamino acids that are biologically active or constitute building blocks for pharmaceutical molecules. Many L-TAs, showing different stereospecificity towards the Cβ configuration, have been isolated. Because of their potential to carry out diastereoselective syntheses, L-TAs have been subjected to structural, functional and mechanistic studies. Nevertheless, their catalytic mechanism and the structural bases of their stereospecificity have not been elucidated. In this study, we have determined the crystal structure of low-specificity L-TA from Escherichia coli at 2.2-Å resolution, in the unliganded form and cocrystallized with L-serine and L-threonine. Furthermore, several active site mutants have been functionally characterized in order to elucidate the reaction mechanism and the molecular bases of stereospecificity. No active site catalytic residue was revealed, and a structural water molecule was assumed to act as the catalytic base in the retro-aldol cleavage reaction. Interestingly, the very large active site opening of E. coli L-TA suggests that much larger molecules than L-threonine isomers may be easily accommodated, and L-TAs may actually have diverse physiological functions in different organisms. Substrate recognition and reaction specificity seem to be guided by the overall microenvironment that surrounds the substrate at the enzyme active site, rather than by one ore more specific residues.
Organizational Affiliation:
Dipartimento di Scienze Biochimiche 'A. Rossi Fanelli', Sapienza Università di Roma, Italy.