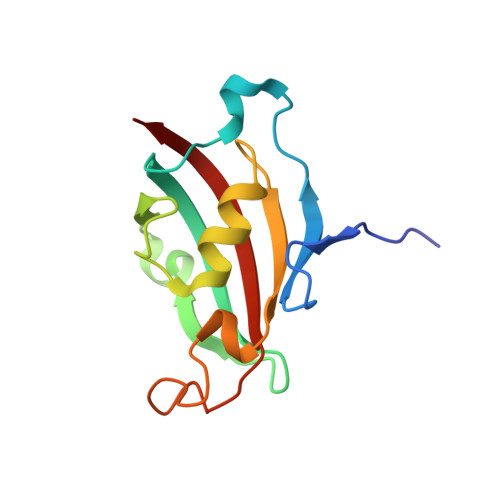
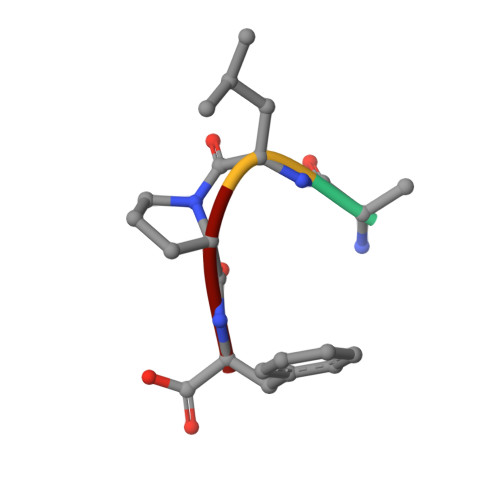
Structural insights into substrate binding by PvFKBP35, a peptidylprolyl cis-trans isomerase from the human malarial parasite Plasmodium vivax
Alag, R., Balakrishna, A.M., Rajan, S., Qureshi, I.A., Shin, J., Lescar, J., Gruber, G., Yoon, H.S.(2013) Eukaryot Cell 12: 627-634
- PubMed: 23435727
- DOI: https://doi.org/10.1128/EC.00016-13
- Primary Citation of Related Structures:
3NI6, 3PA7, 4ITZ - PubMed Abstract:
The immunosuppressive drug FK506 binding proteins (FKBPs), an immunophilin family with the immunosuppressive drug FK506 binding property, exhibit peptidylprolyl cis-trans isomerase (PPIase) activity. While the cyclophilin-catalyzed peptidylprolyl isomerization of X-Pro peptide bonds has been extensively studied, the mechanism of the FKBP-mediated peptidylprolyl isomerization remains uncharacterized. Thus, to investigate the binding of FKBP with its substrate and the underlying catalytic mechanism of the FKBP-mediated proline isomerization, here we employed the FK506 binding domain (FKBD) of the human malarial parasite Plasmodium vivax FK506 binding protein 35 (PvFKBP35) and examined the details of the molecular interaction between the isomerase and a peptide substrate. The crystallographic structures of apo PvFKBD35 and its complex with the tetrapeptide substrate succinyl-Ala-Leu-Pro-Phe-p-nitroanilide (sALPFp) determined at 1.4 Å and 1.65 Å resolutions, respectively, showed that the substrate binds to PvFKBD35 in a cis conformation. Nuclear magnetic resonance (NMR) studies demonstrated the chemical shift perturbations of D55, H67, V73, and I74 residues upon the substrate binding. In addition, the X-ray crystal structure, along with the mutational studies, shows that Y100 is a key residue for the catalytic activity. Taken together, our results provide insights into the catalytic mechanism of PvFKBP35-mediated cis-trans isomerization of substrate and ultimately might aid designing substrate mimetic inhibitors targeting the malarial parasite FKBPs.
Organizational Affiliation:
School of Biological Sciences, Nanyang Technological University, Singapore.