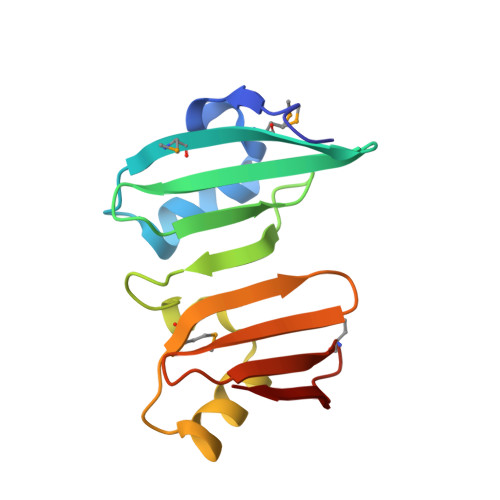
The structure of BVU2987 from Bacteroides vulgatus reveals a superfamily of bacterial periplasmic proteins with possible inhibitory function.
Das, D., Finn, R.D., Carlton, D., Miller, M.D., Abdubek, P., Astakhova, T., Axelrod, H.L., Bakolitsa, C., Chen, C., Chiu, H.J., Chiu, M., Clayton, T., Deller, M.C., Duan, L., Ellrott, K., Ernst, D., Farr, C.L., Feuerhelm, J., Grant, J.C., Grzechnik, A., Han, G.W., Jaroszewski, L., Jin, K.K., Klock, H.E., Knuth, M.W., Kozbial, P., Krishna, S.S., Kumar, A., Marciano, D., McMullan, D., Morse, A.T., Nigoghossian, E., Nopakun, A., Okach, L., Puckett, C., Reyes, R., Rife, C.L., Sefcovic, N., Tien, H.J., Trame, C.B., van den Bedem, H., Weekes, D., Wooten, T., Xu, Q., Hodgson, K.O., Wooley, J., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 1265-1273
- PubMed: 20944221
- DOI: https://doi.org/10.1107/S1744309109046788
- Primary Citation of Related Structures:
3DUE - PubMed Abstract:
Proteins that contain the DUF2874 domain constitute a new Pfam family PF11396. Members of this family have predominantly been identified in microbes found in the human gut and oral cavity. The crystal structure of one member of this family, BVU2987 from Bacteroides vulgatus, has been determined, revealing a β-lactamase inhibitor protein-like structure with a tandem repeat of domains. Sequence analysis and structural comparisons reveal that BVU2987 and other DUF2874 proteins are related to β-lactamase inhibitor protein, PepSY and SmpA_OmlA proteins and hence are likely to function as inhibitory proteins.
Organizational Affiliation:
Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, Menlo Park, CA, USA.