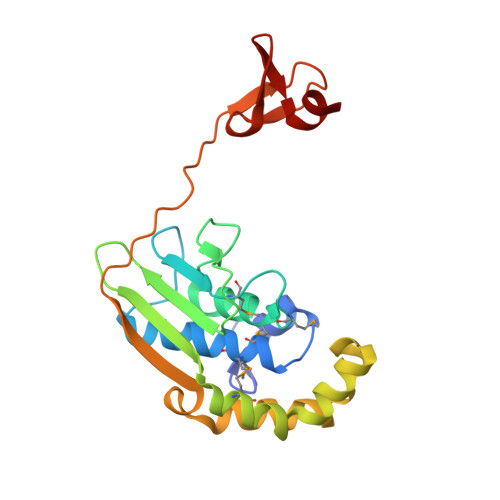
Structures of three members of Pfam PF02663 (FmdE) implicated in microbial methanogenesis reveal a conserved alpha+beta core domain and an auxiliary C-terminal treble-clef zinc finger.
Axelrod, H.L., Das, D., Abdubek, P., Astakhova, T., Bakolitsa, C., Carlton, D., Chen, C., Chiu, H.J., Clayton, T., Deller, M.C., Duan, L., Ellrott, K., Farr, C.L., Feuerhelm, J., Grant, J.C., Grzechnik, A., Han, G.W., Jaroszewski, L., Jin, K.K., Klock, H.E., Knuth, M.W., Kozbial, P., Krishna, S.S., Kumar, A., Lam, W.W., Marciano, D., McMullan, D., Miller, M.D., Morse, A.T., Nigoghossian, E., Nopakun, A., Okach, L., Puckett, C., Reyes, R., Sefcovic, N., Tien, H.J., Trame, C.B., van den Bedem, H., Weekes, D., Wooten, T., Xu, Q., Hodgson, K.O., Wooley, J., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 1335-1346
- PubMed: 20944230
- DOI: https://doi.org/10.1107/S1744309110020166
- Primary Citation of Related Structures:
2GLZ, 2GVI, 3D00 - PubMed Abstract:
Examination of the genomic context for members of the FmdE Pfam family (PF02663), such as the protein encoded by the fmdE gene from the methanogenic archaeon Methanobacterium thermoautotrophicum, indicates that 13 of them are co-transcribed with genes encoding subunits of molybdenum formylmethanofuran dehydrogenase (EC 1.2.99.5), an enzyme that is involved in microbial methane production. Here, the first crystal structures from PF02663 are described, representing two bacterial and one archaeal species: B8FYU2_DESHY from the anaerobic dehalogenating bacterium Desulfitobacterium hafniense DCB-2, Q2LQ23_SYNAS from the syntrophic bacterium Syntrophus aciditrophicus SB and Q9HJ63_THEAC from the thermoacidophilic archaeon Thermoplasma acidophilum. Two of these proteins, Q9HJ63_THEAC and Q2LQ23_SYNAS, contain two domains: an N-terminal thioredoxin-like α+β core domain (NTD) consisting of a five-stranded, mixed β-sheet flanked by several α-helices and a C-terminal zinc-finger domain (CTD). B8FYU2_DESHY, on the other hand, is composed solely of the NTD. The CTD of Q9HJ63_THEAC and Q2LQ23_SYNAS is best characterized as a treble-clef zinc finger. Two significant structural differences between Q9HJ63_THEAC and Q2LQ23_SYNAS involve their metal binding. First, zinc is bound to the putative active site on the NTD of Q9HJ63_THEAC, but is absent from the NTD of Q2LQ23_SYNAS. Second, whereas the structure of the CTD of Q2LQ23_SYNAS shows four Cys side chains within coordination distance of the Zn atom, the structure of Q9HJ63_THEAC is atypical for a treble-cleft zinc finger in that three Cys side chains and an Asp side chain are within coordination distance of the zinc.
Organizational Affiliation:
Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, Menlo Park, CA, USA.