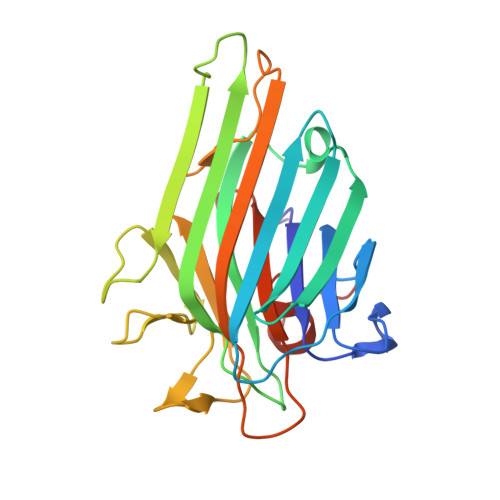
Crystal structures of Cratylia floribunda seed lectin at acidic and basic pHs. Insights into the structural basis of the pH-dependent dimer-tetramer transition.
Del Sol, F.G., Cavada, B.S., Calvete, J.J.(2007) J Struct Biol 158: 1-9
- PubMed: 17251039
- DOI: https://doi.org/10.1016/j.jsb.2006.08.014
- Primary Citation of Related Structures:
2D3P, 2D3R - PubMed Abstract:
Structural determinants underlaying the pH-dependent dimer-tetramer transition of Diocleinae lectins were investigated from the structures of Cratylia floribunda seed lectin crystallized in conditions where it exist as a dimer (pH 4.6) or as a tetramer (pH 8.5). The acidic (aCFL) and the basic (bCFL) tetramers superimpose with overall r.m.s.d. of 0.53 A, though interdimer contacts are drastically reduced in aCFL, and the r.m.s.d. for the superposition of the 117-120 loops of aCFL vs. the bCFL tetramer is 1.29 A. Our data support the view that His51 plays a role in determining the conformation of the central cavity loops and that interdimer contacts involving ordered loop residues stabilize the canonical, pH-dependent tetramer. In the bCFL tetramer, hydrogen bonds between Asn118 and Thr120 of monomers A and D and residues Ser66, Ser108, Ser110, and Thr49 of the opposite monomer stabilize the canonical, pH-dependent tetrameric lectin structure. In CFL, Asn131 makes intradimer contacts with Asn122 and Ala123. In comparison, His131 in Dioclea grandiflora lectin establishes a network of interdimer interactions bridging the four central loops of the pH-independent tetramer. Our data provide new insights into the participation of specific amino acid residues in the mechanism of the quaternary association of Diocleinae lectins.
Organizational Affiliation:
Instituto de Biomedicina de Valencia, C.S.I.C., E-46010 Valencia, Spain.