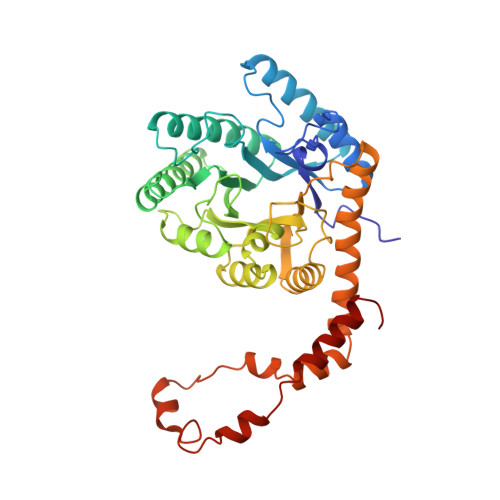
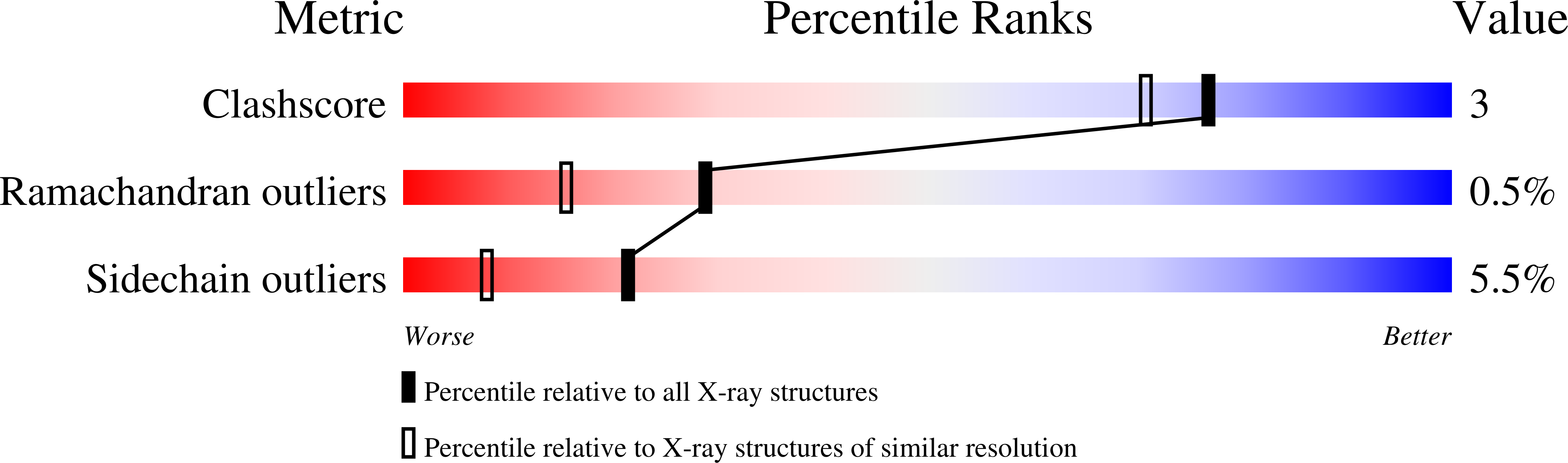X-ray crystallographic structures of D-xylose isomerase-substrate complexes position the substrate and provide evidence for metal movement during catalysis.
Lavie, A., Allen, K.N., Petsko, G.A., Ringe, D.(1994) Biochemistry 33: 5469-5480
- PubMed: 8180169
- DOI: https://doi.org/10.1021/bi00184a016
- Primary Citation of Related Structures:
1XYA, 1XYB, 1XYC - PubMed Abstract:
The X-ray crystallographic structures of the metal-activated enzyme xylose isomerase from Streptomyces olivochromogenes with the substrates D-glucose, 3-O-methyl-D-glucose and in the absence of substrate were determined to 1.96-, 2.19-, and 1.81-A resolution and refined to R-factors of 16.6%, 15.9%, and 16.1%, respectively. Xylose isomerase catalyzes the interconversion between glucose and fructose (xylose and xylulose under physiological conditions) by utilizing two metal cofactors to promote a hydride shift; the metals are bridged by a glutamate residue. This puts xylose isomerase in the small but rapidly growing family of enzymes with a bridged bimetallic active site, in which both metals are involved in the chemical transformation. The substrate 3-O-methylglucose was chosen in order to position the glucose molecule in the observed electron density unambiguously. Of the two essential magnesium ions per active site, Mg-2 was observed to occupy two alternate positions, separated by 1.8 A, in the substrate-soaked structures. The deduced movement was not observed in the structure without substrate present and is attributed to a step following substrate binding but prior to isomerization. The substrates glucose and 3-O-methylglucose are observed in their linear extended forms and make identical interactions with the enzyme by forming ligands to Mg-1 through O2 and O4 and by forming hydrogen bonds with His53 through O5 and Lys182 through O1. Mg-2 has a water ligand that is interpreted in the crystal structure in the absence of substrate as a hydroxide ion and in the presence of substrate as a water molecule. This hydroxide ion may act as a base to deprotonate the glucose O2 and subsequently protonate the product fructose O1 concomitant with hydride transfer. Calculations of the solvent-accessible surface of possible dimers, with and without the alpha-helical C-terminal domain, suggest that the tetramer is the active form of this xylose isomerase.
Organizational Affiliation:
Department of Biochemistry, Brandeis University, Massachusetts 02254-9110.