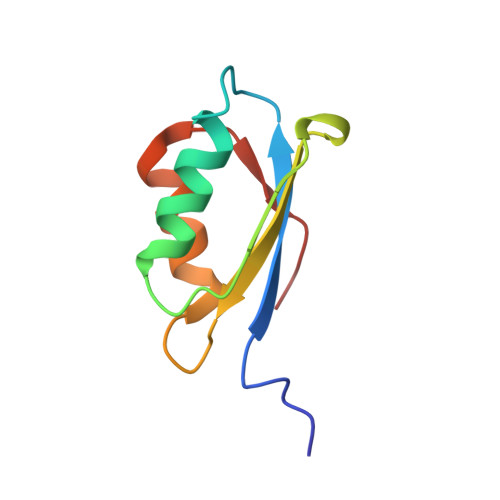
Solution structures of the reduced and Cu(I) bound forms of the first metal binding sequence of ATP7A associated with Menkes disease.
DeSilva, T.M., Veglia, G., Opella, S.J.(2005) Proteins 61: 1038-1049
- PubMed: 16211579
- DOI: https://doi.org/10.1002/prot.20639
- Primary Citation of Related Structures:
1KVI, 1KVJ - PubMed Abstract:
The coding sequence for the first N-terminal copper binding motif of the human Menkes disease protein (MNK1; residues 2-79) was synthesized, cloned, and expressed in bacteria for biochemical and structural studies. MNK1 adopts the betaalphabetabetaalphabeta fold common to all the metal binding sequences (MBS) found in other metal transport systems (e.g., the yeast copper chaperone for superoxide dismutase CCS, the yeast copper chaperone ATX1 bound to Hg(II), and most recently Cu(I), the bacterial copper binding protein, CopZ, and the bacterial Hg(II) binding protein MerP), although substantial differences were found in the metal binding loop. Similar to ATX1, MNK1 binds Cu(I) in a distorted linear bicoordinate geometry. As with MerP, MNK1 has a high affinity for both Hg(II) and Cu(I), although it displays a marked preference for Cu(I). In addition, we found that F71 is a key residue in the compact folding of MNK1, and its mutation to alanine results in an unfolded structure. The homologous residue in MerP has also been mutated with similar results. Finally, to understand the relationship between protein folding and metal affinity and specificity, we expressed a chimeric MBS with the MNK1 protein carrying the binding motif of MerP (CAAC-MNK1); this chimeric protein showed differences in structure and the dynamics of the binding site that may account for metal specificity.
Organizational Affiliation:
Department of Neurology, Children's Hospital, Harvard Medical School, Boston, Massachusetts, USA.