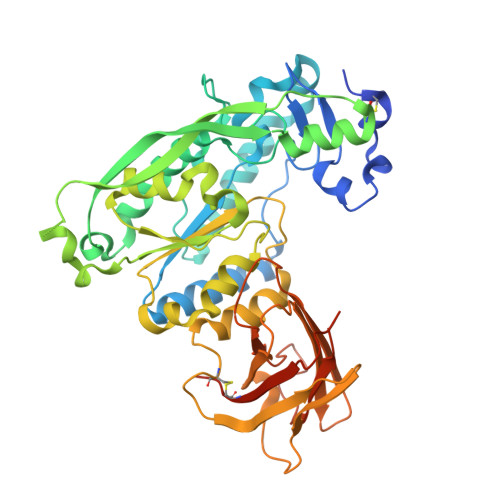
Structural and Functional Characterization of Pseudomonas aeruginosa AlgX: ROLE OF AlgX IN ALGINATE ACETYLATION.
Riley, L.M., Weadge, J.T., Baker, P., Robinson, H., Codee, J.D., Tipton, P.A., Ohman, D.E., Howell, P.L.(2013) J Biol Chem 288: 22299-22314
- PubMed: 23779107
- DOI: https://doi.org/10.1074/jbc.M113.484931
- Primary Citation of Related Structures:
4KNC - PubMed Abstract:
The exopolysaccharide alginate, produced by mucoid Pseudomonas aeruginosa in the lungs of cystic fibrosis patients, undergoes two different chemical modifications as it is synthesized that alter the properties of the polymer and hence the biofilm. One modification, acetylation, causes the cells in the biofilm to adhere better to lung epithelium, form microcolonies, and resist the effects of the host immune system and/or antibiotics. Alginate biosynthesis requires 12 proteins encoded by the algD operon, including AlgX, and although this protein is essential for polymer production, its exact role is unknown. In this study, we present the X-ray crystal structure of AlgX at 2.15 Å resolution. The structure reveals that AlgX is a two-domain protein, with an N-terminal domain with structural homology to members of the SGNH hydrolase superfamily and a C-terminal carbohydrate-binding module. A number of residues in the carbohydrate-binding module form a substrate recognition "pinch point" that we propose aids in alginate binding and orientation. Although the topology of the N-terminal domain deviates from canonical SGNH hydrolases, the residues that constitute the Ser-His-Asp catalytic triad characteristic of this family are structurally conserved. In vivo studies reveal that site-specific mutation of these residues results in non-acetylated alginate. This catalytic triad is also required for acetylesterase activity in vitro. Our data suggest that not only does AlgX protect the polymer as it passages through the periplasm but that it also plays a role in alginate acetylation. Our results provide the first structural insight for a wide group of closely related bacterial polysaccharide acetyltransferases.
Organizational Affiliation:
Program in Molecular Structure and Function, The Hospital for Sick Children, Toronto, Ontario M5G 1X8, Canada.