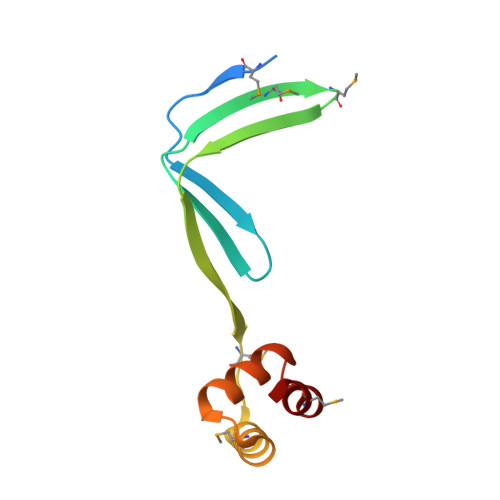
The crystal structure of archaeal nascent polypeptide-associated complex (NAC) reveals a unique fold and the presence of a UBA domain
Spreter, T., Pech, M., Beatrix, B.(2005) J Biol Chem 280: 15849-15854
- PubMed: 15665334
- DOI: https://doi.org/10.1074/jbc.M500160200
- Primary Citation of Related Structures:
1TR8 - PubMed Abstract:
Nascent polypeptide-associated complex (NAC) was identified in eukaryotes as the first cytosolic factor that contacts the nascent polypeptide chain emerging from the ribosome. NAC is highly conserved from yeast to humans. Mutations in NAC cause severe embryonically lethal phenotypes in mice, Drosophila, and Caenorhabditis elegans. NAC was suggested to protect the nascent chain from inappropriate early interactions with cytosolic factors. Eukaryotic NAC is a heterodimer with two subunits sharing substantial homology with each other. All sequenced archaebacterial genomes exhibit only one gene homologous to the NAC subunits. Here we present the first archaebacterial NAC homolog. It forms a homodimer, and as eukaryotic NAC it is associated with ribosomes and contacts the emerging nascent chain on the ribosome. We present the first crystal structure of a NAC protein revealing two structural features: (i) a novel unique protein fold that mediates dimerization of the complex, and (ii) a ubiquitin-associated domain that suggests a yet unidentified role for NAC in the cellular protein quality control system via the ubiquitination pathway. Based on the presented structure we propose a model for the eukaryotic heterodimeric NAC domain.
Organizational Affiliation:
Institute for Chemistry-Crystallography, Free University of Berlin, Takustrasse 6, D-14195 Berlin, Germany.