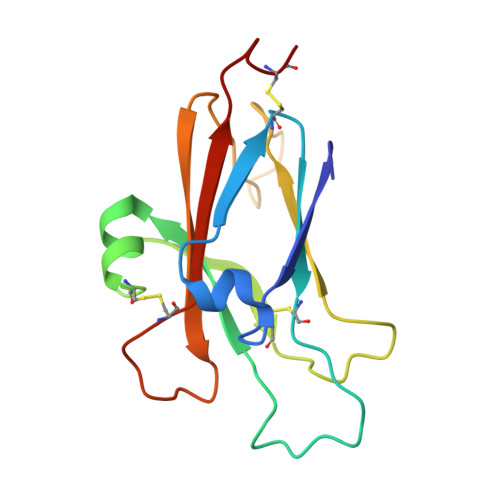
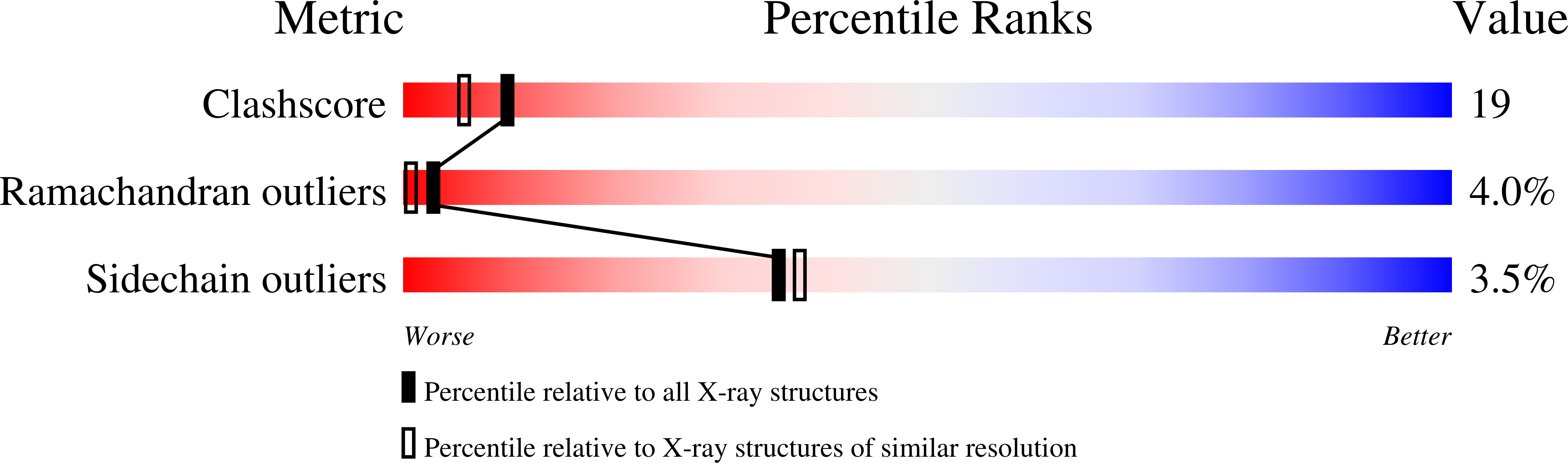Three distinct molecular surfaces in ephrin-A5 are essential for a functional interaction with EphA3.
Day, B., To, C., Himanen, J.P., Smith, F.M., Nikolov, D.B., Boyd, A.W., Lackmann, M.(2005) J Biol Chem 280: 26526-26532
- PubMed: 15901737
- DOI: https://doi.org/10.1074/jbc.M504972200
- Primary Citation of Related Structures:
1SHX - PubMed Abstract:
Eph receptor tyrosine kinases (Ephs) function as molecular relays that interact with cell surface-bound ephrin ligands to direct the position of migrating cells. Structural studies revealed that, through two distinct contact surfaces on opposite sites of each protein, Eph and ephrin binding domains assemble into symmetric, circular heterotetramers. However, Eph signal initiation requires the assembly of higher order oligomers, suggesting additional points of contact. By screening a random library of EphA3 binding-compromised ephrin-A5 mutants, we have now determined ephrin-A5 residues that are essential for the assembly of high affinity EphA3 signaling complexes. In addition to the two interfaces predicted from the crystal structure of the homologous EphB2.ephrin-B2 complex, we identified a cluster of 10 residues on the ephrin-A5 E alpha-helix, the E-F loop, the underlying H beta-strand, as well as the nearby B-C loop, which define a distinct third surface required for oligomerization and activation of EphA3 signaling. Together with a corresponding third surface region identified recently outside of the minimal ephrin binding domain of EphA3, our findings provide experimental evidence for the essential contribution of three distinct protein-interaction interfaces to assemble functional EphA3 signaling complexes.
Organizational Affiliation:
Queensland Institute of Medical Research, P. O. Royal Brisbane Hospital 4029, Queensland, Australia.