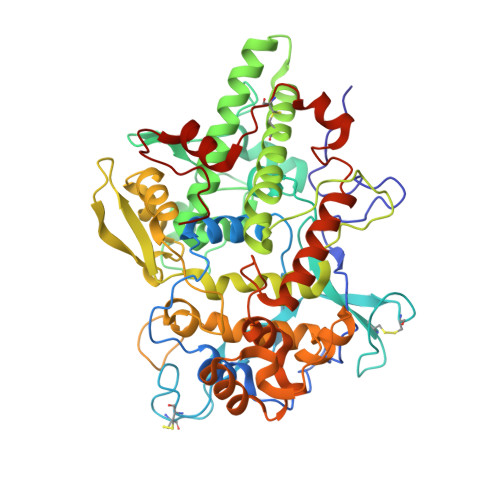
Secreted heme peroxidase from Dictyostelium discoideum: Insights into catalysis, structure, and biological role.
Nicolussi, A., Dunn, J.D., Mlynek, G., Bellei, M., Zamocky, M., Battistuzzi, G., Djinovic-Carugo, K., Furtmuller, P.G., Soldati, T., Obinger, C.(2018) J Biol Chem 293: 1330-1345
- PubMed: 29242189
- DOI: https://doi.org/10.1074/jbc.RA117.000463
- Primary Citation of Related Structures:
6ERC - PubMed Abstract:
Oxidation of halides and thiocyanate by heme peroxidases to antimicrobial oxidants is an important cornerstone in the innate immune system of mammals. Interestingly, phylogenetic and physiological studies suggest that homologous peroxidases are already present in mycetozoan eukaryotes such as Dictyostelium discoideum This social amoeba kills bacteria via phagocytosis for nutrient acquisition at its single-cell stage and for antibacterial defense at its multicellular stages. Here, we demonstrate that peroxidase A from D. discoideum (DdPoxA) is a stable, monomeric, glycosylated, and secreted heme peroxidase with homology to mammalian peroxidases. The first crystal structure (2.5 Å resolution) of a mycetozoan peroxidase of this superfamily shows the presence of a post-translationally-modified heme with one single covalent ester bond between the 1-methyl heme substituent and Glu-236. The metalloprotein follows the halogenation cycle, whereby compound I oxidizes iodide and thiocyanate at high rates (>10 8 m -1 s -1 ) and bromide at very low rates. It is demonstrated that DdPoxA is up-regulated and likely secreted at late multicellular development stages of D. discoideum when migrating slugs differentiate into fruiting bodies that contain persistent spores on top of a cellular stalk. Expression of DdPoxA is shown to restrict bacterial contamination of fruiting bodies. Structure and function of DdPoxA are compared with evolutionary-related mammalian peroxidases in the context of non-specific immune defense.
Organizational Affiliation:
From the Department of Chemistry, Division of Biochemistry, BOKU-University of Natural Resources and Life Sciences, 1190 Vienna, Austria.