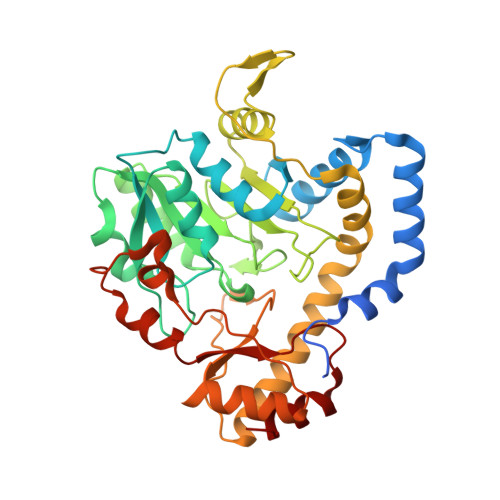
Investigation of the enzymes required for the biosynthesis of 2,3-diacetamido-2,3-dideoxy-d-glucuronic acid in Psychrobacter cryohalolentis K5 T.
Hofmeister, D.L., Seltzner, C.A., Bockhaus, N.J., Thoden, J.B., Holden, H.M.(2023) Protein Sci 32: e4502-e4502
- PubMed: 36346293
- DOI: https://doi.org/10.1002/pro.4502
- Primary Citation of Related Structures:
8E62, 8E75, 8E77 - PubMed Abstract:
Psychrobacter cryohalolentis K5 T is a Gram-negative bacterium first isolated from Siberian permafrost in 2006. It has a complex O-antigen containing l-rhamnose, d-galactose, two diacetamido-sugars, and one triacetamido-sugar. The biosynthetic pathway for one of the diacetamido-sugars, namely 2,3-diacetamido-2,3-dideoxy-d-glucuronic acid, is presently unknown. Utilizing the published genome sequence of P. cryohalolentis K5 T , we hypothesized that the genes designated Pcryo_0613, Pcryo_0614, Pcryo_0616, and Pcryo_0615 encode for a uridine dinucleotide (UDP)-N-acetyl-d-glucosamine 6-dehydrogenase, an nicotinamide adenine dinucleotide (oxidized) (NAD + )-dependent dehydrogenase, a pyridoxal 5'-phosphate (PLP)-dependent aminotransferase, and an N-acetyltransferase, respectively, activities of which would be required for the biosynthesis of this unusual carbohydrate. Here we present the cloning, overexpression, and purification of these hypothetical proteins. Kinetic data on the enzymes encoded by Pcryo_0613, Pcryo_0614, and Pcryo_0615 confirmed their postulated biochemical activities. In addition, the high-resolution X-ray structures of both the internal and external aldimine forms of the aminotransferase were determined to 1.25 and 1.0 Å, respectively. Finally, the three-dimensional architecture of the N-acetyltransferase in complex with its substrate and coenzyme A was solved to 1.8 Å resolution. Strikingly, the N-acetyltransferase was shown to adopt a new motif for UDP-sugar binding. The data presented herein provide additional insight into sugar biosynthesis in Gram-negative bacteria.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin, Madison, Wisconsin, USA.