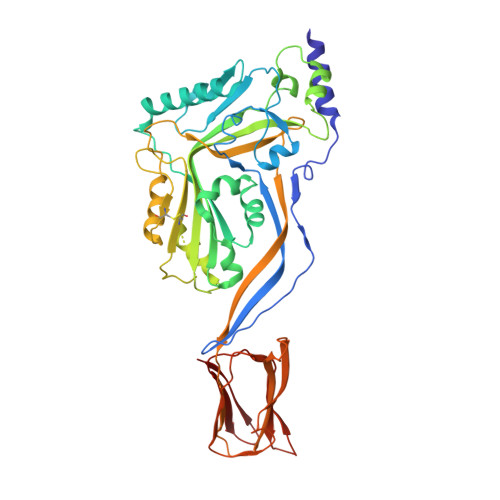
Structural Basis for Receptor Recognition by the Human CD59-Responsive Cholesterol-Dependent Cytolysins.
Lawrence, S.L., Gorman, M.A., Feil, S.C., Mulhern, T.D., Kuiper, M.J., Ratner, A.J., Tweten, R.K., Morton, C.J., Parker, M.W.(2016) Structure 24: 1488-1498
- PubMed: 27499440
- DOI: https://doi.org/10.1016/j.str.2016.06.017
- Primary Citation of Related Structures:
5IMT, 5IMW, 5IMY - PubMed Abstract:
Cholesterol-dependent cytolysins (CDCs) are a family of pore-forming toxins that punch holes in the outer membrane of eukaryotic cells. Cholesterol serves as the receptor, but a subclass of CDCs first binds to human CD59. Here we describe the crystal structures of vaginolysin and intermedilysin complexed to CD59. These studies, together with small-angle X-ray scattering, reveal that CD59 binds to each at different, though overlapping, sites, consistent with molecular dynamics simulations and binding studies. The CDC consensus undecapeptide motif, which for the CD59-responsive CDCs has a proline instead of a tryptophan in the motif, adopts a strikingly different conformation between the structures; our data suggest that the proline acts as a selectivity switch to ensure CD59-dependent CDCs bind their protein receptor first in preference to cholesterol. The structural data suggest a detailed model of how these water-soluble toxins assemble as prepores on the cell surface.
Organizational Affiliation:
ACRF Rational Drug Discovery Centre, St. Vincent's Institute of Medical Research, Fitzroy, VIC 3065, Australia.