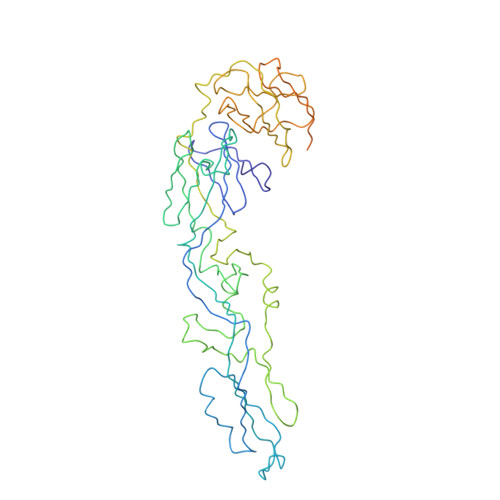
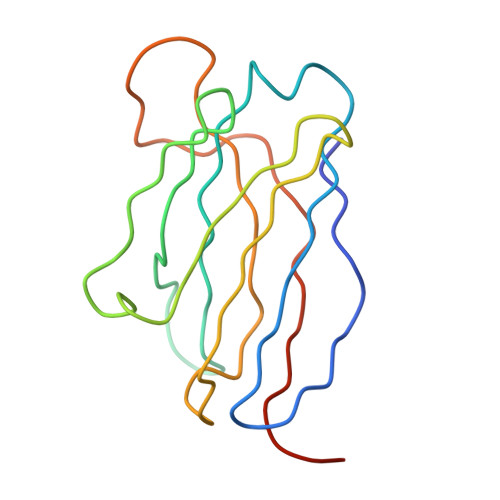
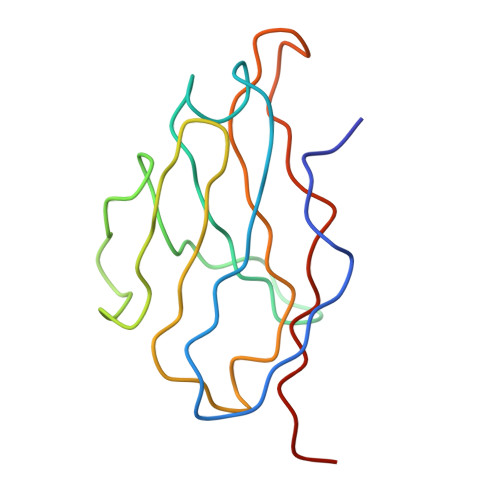
Cryo-EM structure of an antibody that neutralizes dengue virus type 2 by locking E protein dimers.
Fibriansah, G., Ibarra, K.D., Ng, T.S., Smith, S.A., Tan, J.L., Lim, X.N., Ooi, J.S., Kostyuchenko, V.A., Wang, J., de Silva, A.M., Harris, E., Crowe, J.E., Lok, S.M.(2015) Science 349: 88-91
- PubMed: 26138979
- DOI: https://doi.org/10.1126/science.aaa8651
- Primary Citation of Related Structures:
4UIF, 4UIH, 5A1Z - PubMed Abstract:
There are four closely-related dengue virus (DENV) serotypes. Infection with one serotype generates antibodies that may cross-react and enhance infection with other serotypes in a secondary infection. We demonstrated that DENV serotype 2 (DENV2)-specific human monoclonal antibody (HMAb) 2D22 is therapeutic in a mouse model of antibody-enhanced severe dengue disease. We determined the cryo-electron microscopy (cryo-EM) structures of HMAb 2D22 complexed with two different DENV2 strains. HMAb 2D22 binds across viral envelope (E) proteins in the dimeric structure, which probably blocks the E protein reorganization required for virus fusion. HMAb 2D22 "locks" two-thirds of or all dimers on the virus surface, depending on the strain, but neutralizes these DENV2 strains with equal potency. The epitope defined by HMAb 2D22 is a potential target for vaccines and therapeutics.
Organizational Affiliation:
Program in Emerging Infectious Diseases, Duke-National University of Singapore Graduate Medical School, Singapore. Centre for BioImaging Sciences, National University of Singapore, Singapore.