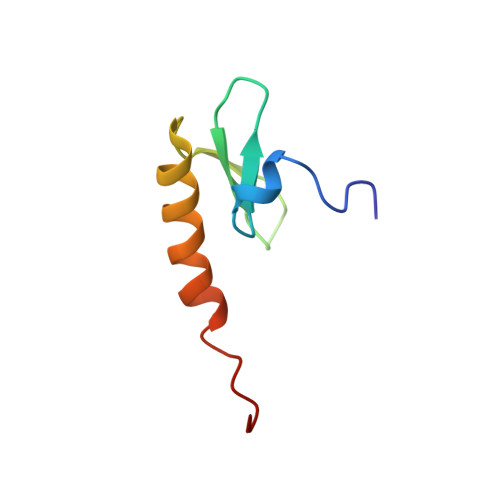
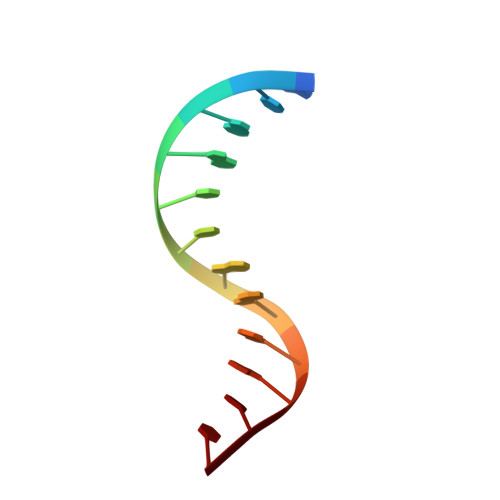
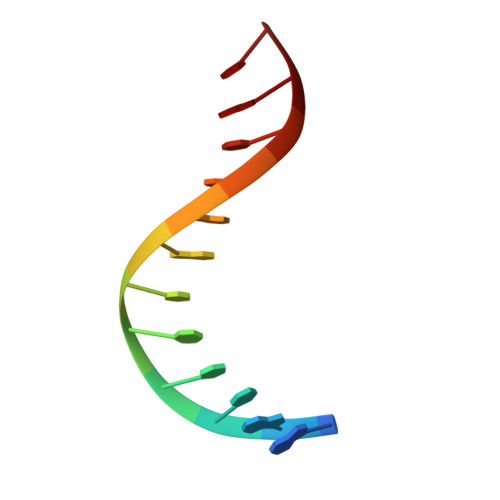
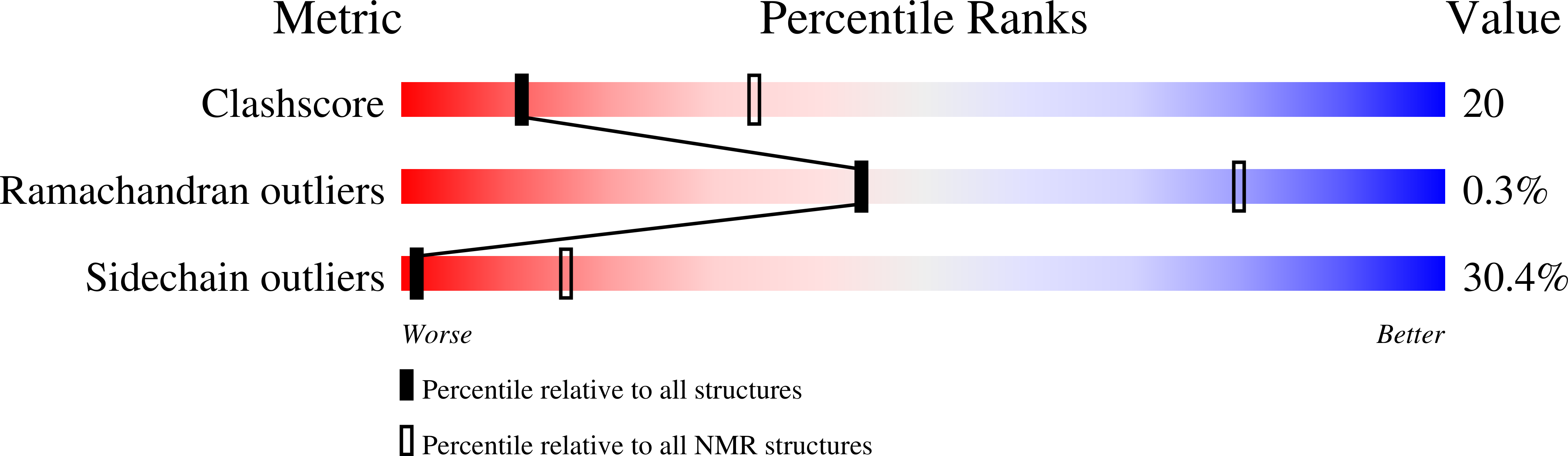NMR Structure of the Amino-Terminal Domain of the Lambda Integrase Protein in Complex with DNA: Immobilization of a Flexible Tail Facilitates Beta- Sheet Recognition of the Major Groove.
Fadeev, E.A., Sam, M.D., Clubb, R.T.(2009) J Mol Biol 388: 682
- PubMed: 19324050
- DOI: https://doi.org/10.1016/j.jmb.2009.03.041
- Primary Citation of Related Structures:
2WCC - PubMed Abstract:
The integrase protein (Int) from bacteriophage lambda is the archetypal member of the tyrosine recombinase family, a large group of enzymes that rearrange DNA in all domains of life. Int catalyzes the insertion and excision of the viral genome into and out of the Escherichia coli chromosome. Recombination transpires within higher-order nucleoprotein complexes that form when its amino-terminal domain binds to arm-type DNA sequences that are located distal to the site of strand exchange. Arm-site binding by Int is essential for catalysis, as it promotes Int-mediated bridge structures that stabilize the recombination machinery. We have elucidated how Int is able to sequence specifically recognize the arm-type site sequence by determining the solution structure of its amino-terminal domain (Int(N), residues Met1 to Leu64) in complex with its P'2 DNA binding site. Previous studies have shown that Int(N) adopts a rare monomeric DNA binding fold that consists of a three-stranded antiparallel beta-sheet that is packed against a carboxy-terminal alpha helix. A low-resolution crystal structure of the full-length protein also revealed that the sheet is inserted into the major groove of the arm-type site. The solution structure presented here reveals how Int(N) specifically recognizes the arm-type site sequence. A novel feature of the new solution structure is the use of an 11-residue tail that is located at the amino terminus. DNA binding induces the folding of a 3(10) helix in the tail that projects the amino terminus of the protein deep into the minor groove for stabilizing DNA contacts. This finding reveals the structural basis for the observation that the "unstructured" amino terminus is required for recombination.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of California, Los Angeles, CA 90095-1570, USA.