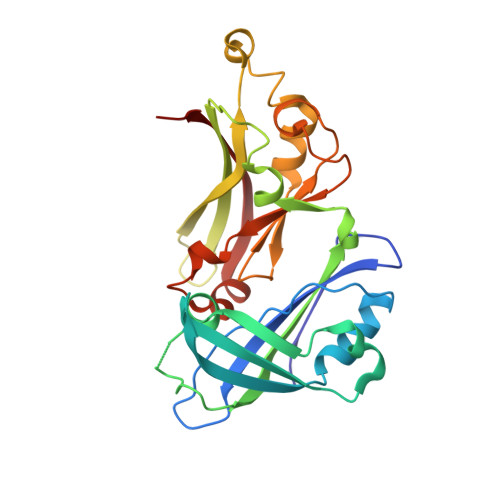
Surface Features of a Mononegavirales Matrix Protein Indicate Sites of Membrane Interaction.
Money, V.A., Mcphee, H.K., Mosely, J.A., Sanderson, J.M., Yeo, R.P.(2009) Proc Natl Acad Sci U S A 106: 4441
- PubMed: 19251668
- DOI: https://doi.org/10.1073/pnas.0805740106
- Primary Citation of Related Structures:
2VQP - PubMed Abstract:
The matrix protein (M) of respiratory syncytial virus (RSV), the prototype viral member of the Pneumovirinae (family Paramyxoviridae, order Mononegavirales), has been crystallized and the structure determined to a resolution of 1.6 A. The structure comprises 2 compact beta-rich domains connected by a relatively unstructured linker region. Due to the high degree of side-chain order in the structure, an extensive contiguous area of positive surface charge covering approximately 600 A(2) can be resolved. This unusually large patch of positive surface potential spans both domains and the linker, and provides a mechanism for driving the interaction of the protein with a negatively-charged membrane surface or other virion components such as the nucleocapsid. This patch is complemented by regions of high hydrophobicity and a striking planar arrangement of tyrosine residues encircling the C-terminal domain. Comparison of the RSV M sequence with other members of the Pneumovirinae shows that regions of divergence correspond to surface exposed loops in the M structure, with the majority of viral species-specific differences occurring in the N-terminal domain.
Organizational Affiliation:
The Maurice Wilkins Centre for Molecular Biodiscovery and the School of Biological Sciences, University of Auckland, Thomas Building, 3a Symonds Street, Auckland Central 1010, New Zealand.