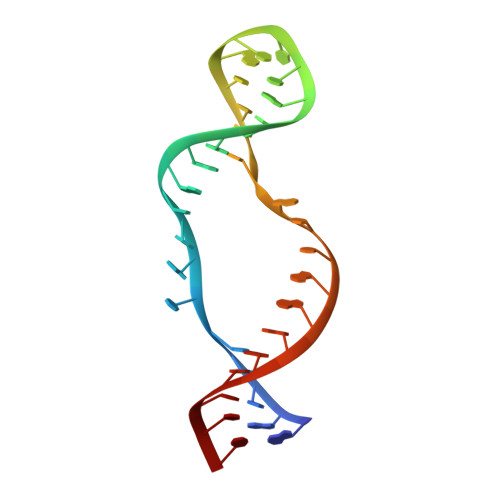
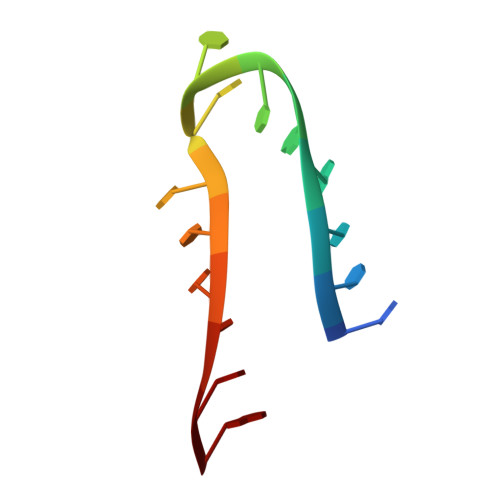H/ACA small nucleolar RNA pseudouridylation pockets bind substrate RNA to form three-way junctions that position the target U for modification.
Wu, H., Feigon, J.(2007) Proc Natl Acad Sci U S A 104: 6655-6660
- PubMed: 17412831
- DOI: https://doi.org/10.1073/pnas.0701534104
- Primary Citation of Related Structures:
2P89 - PubMed Abstract:
During the biogenesis of eukaryotic ribosomal RNA (rRNA) and spliceosomal small nuclear RNA (snRNA), uridines at specific sites are converted to pseudouridines by H/ACA ribonucleoprotein particles (RNPs). Each H/ACA RNP contains a substrate-specific H/ACA RNA and four common proteins, the pseudouridine synthase Cbf5, Nop10, Gar1, and Nhp2. The H/ACA RNA contains at least one pseudouridylation (psi) pocket, which is complementary to the sequences flanking the target uridine. In this article, we show structural evidence that the psi pocket can form the predicted base pairs with substrate RNA in the absence of protein components. We report the solution structure of the complex between an RNA hairpin derived from the 3' psi pocket of human U65 H/ACA small nucleolar RNA (snoRNA) and the substrate rRNA. The snoRNA-rRNA substrate complex has a unique structure with two offset parallel pairs of stacked helices and two unusual intermolecular three-way junctions, which together organize the substrate for docking into the active site of Cbf5. The substrate RNA interacts on one face of the snoRNA in the complex, forming a structure that easily could be accommodated in the H/ACA RNP, and explains how successive substrate RNAs could be loaded onto and unloaded from the H/ACA RNA in the RNP.
Organizational Affiliation:
Department of Chemistry and Biochemistry, and Molecular Biology Institute, University of California, Los Angeles, CA 90095-1569, USA.