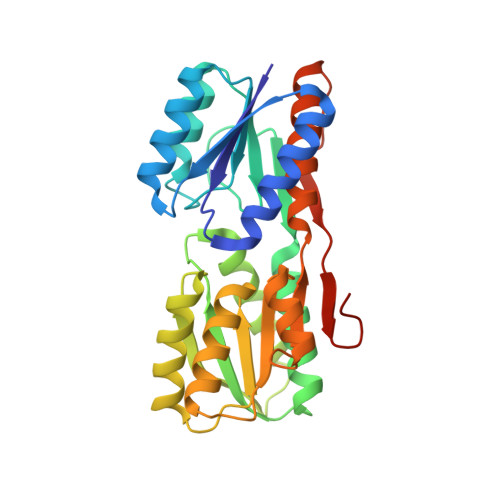
The backbone structure of the thermophilic Thermoanaerobacter tengcongensis ribose binding protein is essentially identical to its mesophilic E. coli homolog.
Cuneo, M.J., Tian, Y., Allert, M., Hellinga, H.W.(2008) BMC Struct Biol 8: 20-20
- PubMed: 18373848
- DOI: https://doi.org/10.1186/1472-6807-8-20
- Primary Citation of Related Structures:
2IOY - PubMed Abstract:
Comparison of experimentally determined mesophilic and thermophilic homologous protein structures is an important tool for understanding the mechanisms that contribute to thermal stability. Of particular interest are pairs of homologous structures that are structurally very similar, but differ significantly in thermal stability. We report the X-ray crystal structure of a Thermoanaerobacter tengcongensis ribose binding protein (tteRBP) determined to 1.9 A resolution. We find that tteRBP is significantly more stable (appTm value approximately 102 degrees C) than the mesophilic Escherichia coli ribose binding protein (ecRBP) (appTm value ~56 degrees C). The tteRBP has essentially the identical backbone conformation (0.41 A RMSD of 235/271 Calpha positions and 0.65 A RMSD of 270/271 Calpha positions) as ecRBP. Classification of the amino acid substitutions as a function of structure therefore allows the identification of amino acids which potentially contribute to the observed thermal stability of tteRBP in the absence of large structural heterogeneities. The near identity of backbone structures of this pair of proteins entails that the significant differences in their thermal stabilities are encoded exclusively by the identity of the amino acid side-chains. Furthermore, the degree of sequence divergence is strongly correlated with structure; with a high degree of conservation in the core progressing to increased diversity in the boundary and surface regions. Different factors that may possibly contribute to thermal stability appear to be differentially encoded in each of these regions of the protein. The tteRBP/ecRBP pair therefore offers an opportunity to dissect contributions to thermal stability by side-chains alone in the absence of large structural differences.
Organizational Affiliation:
The Institute for Biological Structure and Design and the Department of Biochemistry, Duke University Medical Center, Durham, North Carolina, 27710, USA. mjc18@duke.edu