Structural basis of chaperone-subunit complex recognition by the type 1 pilus assembly platform FimD.
Nishiyama, M., Horst, R., Eidam, O., Herrmann, T., Ignatov, O., Vetsch, M., Bettendorff, P., Jelesarov, I., Glockshuber, R., Capitani, G.(2005) EMBO J 24: 2075-2086
- PubMed: 15920478
- DOI: https://doi.org/10.1038/sj.emboj.7600693
- Primary Citation of Related Structures:
1ZDV, 1ZDX, 1ZE3 - PubMed Abstract:
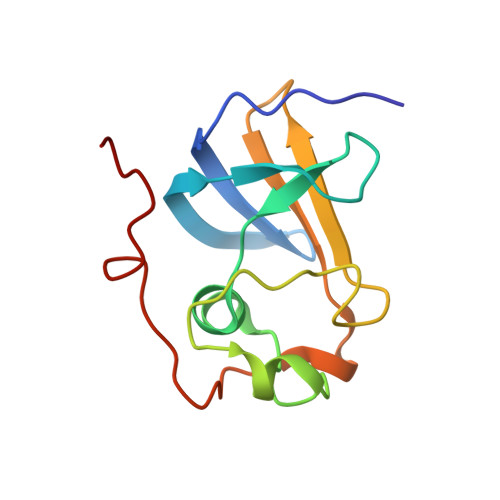
Adhesive type 1 pili from uropathogenic Escherichia coli are filamentous protein complexes that are attached to the assembly platform FimD in the outer membrane. During pilus assembly, FimD binds complexes between the chaperone FimC and type 1 pilus subunits in the periplasm and mediates subunit translocation to the cell surface. Here we report nuclear magnetic resonance and X-ray protein structures of the N-terminal substrate recognition domain of FimD (FimD(N)) before and after binding of a chaperone-subunit complex. FimD(N) consists of a flexible N-terminal segment of 24 residues, a structured core with a novel fold, and a C-terminal hinge segment. In the ternary complex, residues 1-24 of FimD(N) specifically interact with both FimC and the subunit, acting as a sensor for loaded FimC molecules. Together with in vivo complementation studies, we show how this mechanism enables recognition and discrimination of different chaperone-subunit complexes by bacterial pilus assembly platforms.
Organizational Affiliation:
Institut für Molekularbiologie und Biophysik, Eidgenössische Technische Hochschule Hönggerberg, Zürich, Switzerland.