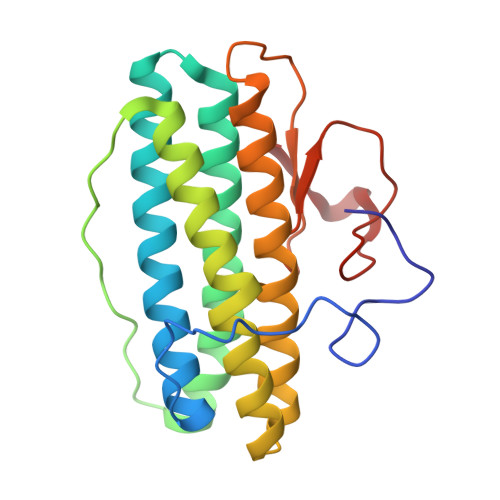
High-resolution crystal structures of Desulfovibrio vulgaris (Hildenborough) nigerythrin: facile, redox-dependent iron movement, domain interface variability, and peroxidase activity in the rubrerythrins.
Iyer, R.B., Silaghi-Dumitrescu, R., Kurtz, D.M., Lanzilotta, W.N.(2005) J Biol Inorg Chem 10: 407-416
- PubMed: 15895271
- DOI: https://doi.org/10.1007/s00775-005-0650-8
- Primary Citation of Related Structures:
1YUX, 1YUZ, 1YV1 - PubMed Abstract:
High-resolution crystal structures of Desulfovibrio vulgaris nigerythrin (DvNgr), a member of the rubrerythrin (Rbr) family, demonstrate an approximately 2-A movement of one iron (Fe1) of the diiron site from a carboxylate to a histidine ligand upon conversion of the mixed-valent ([Fe2(II),Fe1(III)]) to diferrous states, even at cryogenic temperatures. This Glu<-->His ligand "toggling" of one iron, which also occurs in DvRbr, thus, appears to be a characteristic feature of Rbr-type diiron sites. Unique features of DvNgr revealed by these structures include redox-induced flipping of a peptide carbonyl that reversibly forms a hydrogen bond to the histidine ligand to Fe1 of the diiron site, an intra-subunit proximal orientation of the rubredoxin-(Rub)-like and diiron domains, and an electron transfer pathway consisting of six covalent and two hydrogen bonds connecting the Rub-like iron with Fe2 of the diiron site. This pathway can account for DvNgr's relatively rapid peroxidase turnover. The characteristic combination of iron sites together with the redox-dependent iron toggling between protein ligands can account for the selectivity of Rbrs for hydrogen peroxide over dioxygen.
Organizational Affiliation:
Department of Chemistry, University of Georgia, Athens, GA 30602, USA.