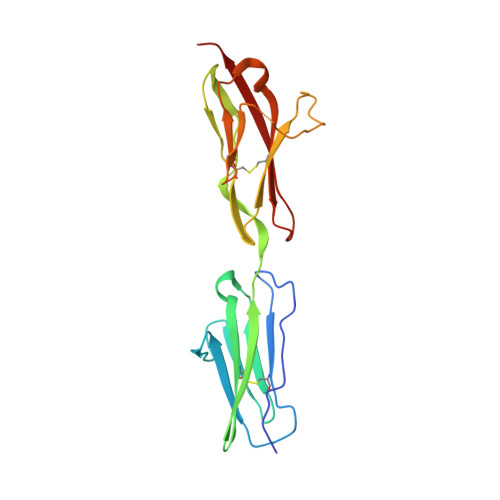
Structural basis for the self-recognition of sDSCAM in Chelicerata.
Cheng, J., Yu, Y., Wang, X., Zheng, X., Liu, T., Hu, D., Jin, Y., Lai, Y., Fu, T.M., Chen, Q.(2023) Nat Commun 14: 2522-2522
- PubMed: 37130844
- DOI: https://doi.org/10.1038/s41467-023-38205-1
- Primary Citation of Related Structures:
7Y4X, 7Y54, 7Y5J, 7Y5R, 7Y6E, 7Y6O, 7Y73, 7Y8H, 7Y8I, 7Y8S, 7Y95, 7Y9A - PubMed Abstract:
To create a functional neural circuit, neurons develop a molecular identity to discriminate self from non-self. The invertebrate Dscam family and vertebrate Pcdh family are implicated in determining synaptic specificity. Recently identified in Chelicerata, a shortened Dscam (sDscam) has been shown to resemble the isoform-generating characters of both Dscam and Pcdh and represent an evolutionary transition. Here we presented the molecular details of sDscam self-recognition via both trans and cis interactions using X-ray crystallographic data and functional assays. Based on our results, we proposed a molecular zipper model for the assemblies of sDscam to mediate cell-cell recognition. In this model, sDscam utilized FNIII domain to form side-by-side interactions with neighboring molecules in the same cell while established hand-in-hand interactions via Ig1 domain with molecules from another cell around. Together, our study provided a framework for understanding the assembly, recognition, and evolution of sDscam.
Organizational Affiliation:
National Clinical Research Center for Geriatrics, West China Hospital, State Key Laboratory of Biotherapy and Collaborative Innovation Center of Biotherapy, Sichuan University, 610041, Chengdu, China.