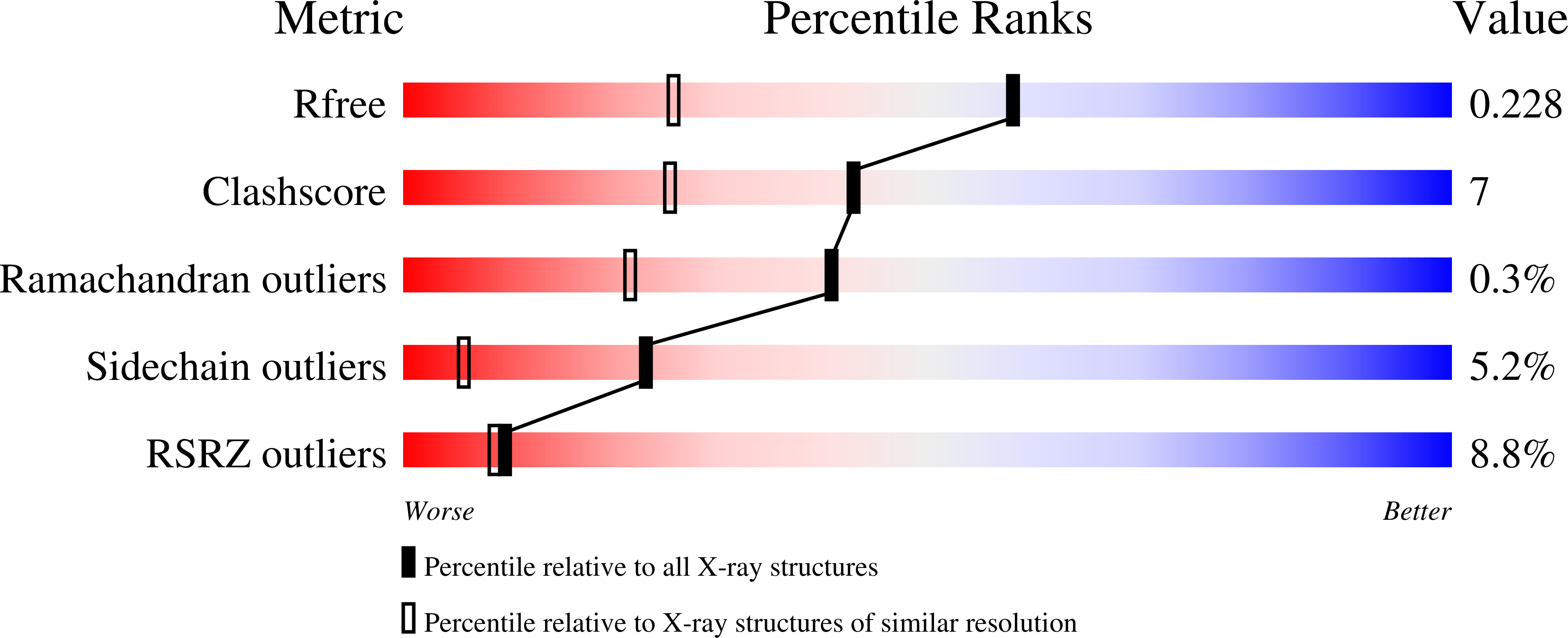
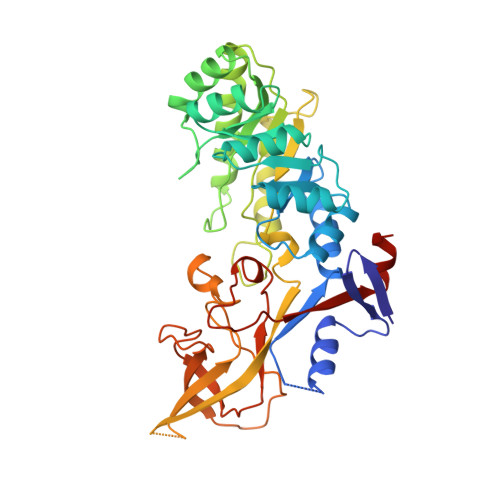
Structure of mammalian ornithine decarboxylase at 1.6 A resolution: stereochemical implications of PLP-dependent amino acid decarboxylases.
Kern, A.D., Oliveira, M.A., Coffino, P., Hackert, M.L.(1999) Structure 7: 567-581
- PubMed: 10378276
- DOI: https://doi.org/10.1016/s0969-2126(99)80073-2
- Primary Citation of Related Structures:
7ODC - PubMed Abstract:
Pyridoxal-5'-phosphate (PLP) dependent enzymes catalyze a broad range of reactions, resulting in bond cleavage at C alpha, C beta, or C gamma carbons of D and L amino acid substrates. Ornithine decarboxylase (ODC) is a PLP-dependent enzyme that controls a critical step in the biosynthesis of polyamines, small organic polycations whose controlled levels are essential for proper growth. ODC inhibition has applications for the treatment of certain cancers and parasitic ailments such as African sleeping sickness.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of Texas at Austin 78712, USA.