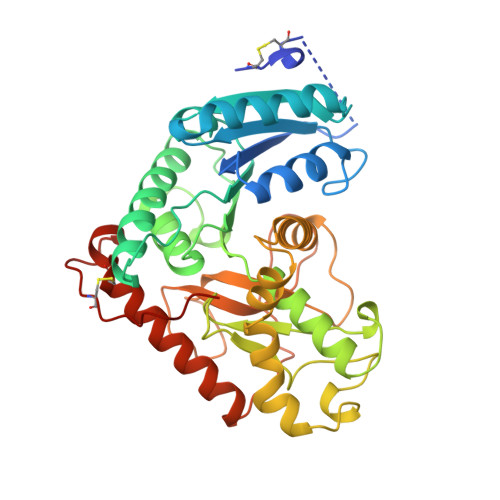
Structural basis for light activation of a chloroplast enzyme: the structure of sorghum NADP-malate dehydrogenase in its oxidized form.
Johansson, K., Ramaswamy, S., Saarinen, M., Lemaire-Chamley, M., Issakidis-Bourguet, E., Miginiac-Maslow, M., Eklund, H.(1999) Biochemistry 38: 4319-4326
- PubMed: 10194350
- DOI: https://doi.org/10.1021/bi982876c
- Primary Citation of Related Structures:
7MDH - PubMed Abstract:
Some key chloroplast enzymes are activated by light via a ferredoxin-thioredoxin reduction system which reduces disulfide bridges in the enzymes. We describe for the first time the structural basis for the redox activation of a chloroplast enzyme, the NADP-dependent malate dehydrogenase (MDH) from Sorghum vulgare whose structure has been determined and refined at 2.4 A resolution. In addition to the normal structural components of MDHs, the enzyme exhibits extensions at both the N- and C-termini, each of which contains a regulatory disulfide bridge which must be reduced for activation. The N-terminal disulfide motif is inserted in a cleft between the two subunits of the dimer, thereby locking the domains in each subunit. The C-terminal disulfide keeps the C-terminal residues tight to the enzyme surface and blocks access to the active site. Reduction of the N-terminal disulfide would release the stopper between the domains and give the enzyme the necessary flexibility. Simultaneous reduction of the C-terminal disulfide would free the C-terminal residues from binding to the enzyme and make the active site accessible.
Organizational Affiliation:
Department of Molecular Biology, Swedish University of Agricultural Sciences, Uppsala.