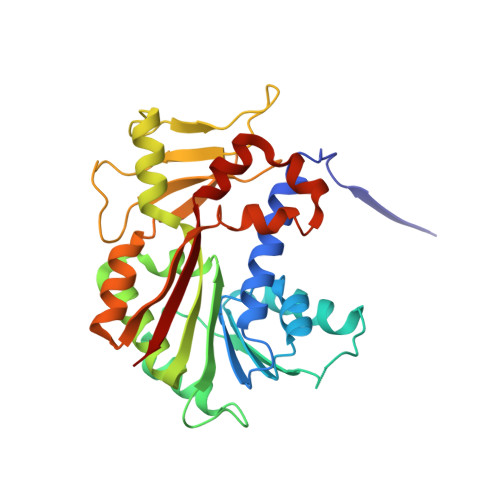
Structure and Biochemical Characteristic of the Methyltransferase (MTase) Domain of RNA Capping Enzyme from African Swine Fever Virus.
Du, X., Gao, Z.Q., Geng, Z., Dong, Y.H., Zhang, H.(2020) J Virol
- PubMed: 33268516
- DOI: https://doi.org/10.1128/JVI.02029-20
- Primary Citation of Related Structures:
7D8U - PubMed Abstract:
African swine fever virus (ASFV) is a complex nucleocytoplasmic large DNA virus (NCLDV) that causes a devastating swine disease and it is urgently needed to develop effective anti-ASFV vaccines and drugs. The process of mRNA 5'-end capping is a common characteristic in eukaryotes and many viruses, and the cap structure is required for mRNA stability and efficient translation. The ASFV protein pNP868R was found to have guanylyltransferase (GTase) activity involved in mRNA capping. Here we report the crystal structure of pNP868R methyltransferase (MTase) domain (referred as pNP868R MT ) in complex with S-adenosyl-L-methionine (AdoMet). The structure shows the characteristic core fold of the class I MTase family and the AdoMet is bound in a negative-deep groove. Remarkably, the N-terminal extension of pNP868R MT is ordered and keeps away from the AdoMet-binding site, distinct from the close conformation over the active site of poxvirus RNA capping D1 subunit or the largely disordered conformation in most cellular RNA capping MTases. Structure-based mutagenesis studies based on the pNP868R MT -cap analog complex model revealed essential residues involved in substrate recognition and binding. Functional studies suggest the N-terminal extension may play an essential role in substrate recognition instead of AdoMet-binding. A positively charged path stretching from the N-terminal extension to the region around the active site was suggested to provide a favorable electrostatic environment for the binding and approaching of substrate RNA into the active site. Our structure and biochemical studies provide novel insights into the methyltransfer process of mRNA cap catalyzed by pNP868R. IMPORTANCE African swine fever (ASF) is a highly contagious hemorrhagic viral disease in pigs that is caused by African swine fever virus (ASFV). There are no effective drugs or vaccines for protection against ASFV infection till now. The protein pNP868R was predicted to be responsible for process of mRNA 5'-end capping in ASFV, which is essential for mRNA stability and efficient translation. Here, we solved the high-resolution crystal structure of the methyltransferase (MTase) domain of pNP868R. The MTase domain structure shows a canonical class I MTase family fold and the AdoMet binds into a negative pocket. Structure-based mutagenesis studies revealed critical and conserved residues involved in AdoMet-binding and substrate RNA-binding. Notably, both the conformation and the role in MTase activities of the N-terminal extension are distinct from those of previously characterized poxvirus MTase domain. Our structure-function studies provide the basis for potential anti-ASFV inhibitor design targeting the critical enzyme.
Organizational Affiliation:
Institute of Health Sciences and School of Life Science, Anhui University, Hefei, Anhui 230601, China.