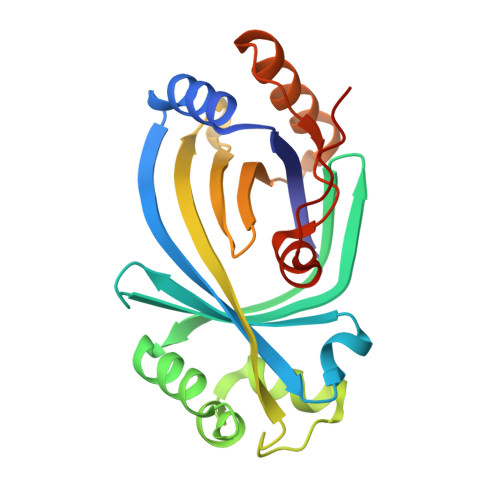
Crystal structure and enzymatic characterization of the putative adenylyl cyclase HpAC1 from Hippeastrum reveal dominant triphosphatase activity.
Kleinboelting, S., Miehling, J., Steegborn, C.(2020) J Struct Biol 212: 107649-107649
- PubMed: 33075486
- DOI: https://doi.org/10.1016/j.jsb.2020.107649
- Primary Citation of Related Structures:
6YP4 - PubMed Abstract:
HpAC1, a protein from Hippeastrum hybrid cultivars, was previously suggested to be a plant adenylyl cyclase. We describe a structural and enzymatic characterization of HpAC1. A crystal structure of HpAC1 in complex with a non-hydrolyzable GTP analog confirms a generic CYTH architecture, comprising a β-barrel with an internal substrate site. The structure reveals significant active site differences to AC proteins with CYTH fold, however, and we find that HpAC1 lacks measurable AC activity. Instead, HpAC1 has substantial triphosphatase activity, indicating this protective activity or a related activity as the protein's physiological function.
Organizational Affiliation:
Department of Biochemistry, University of Bayreuth, Universitaetsstr. 30, 95447 Bayreuth, Germany; Faculty of Chemistry and Chemical Biology, TU Dortmund and Drug Discovery Hub Dortmund (DDHD), Zentrum für Wirkstoffforschung (ZIW), Otto-Hahn-Strasse 4a, 44227 Dortmund, Germany.