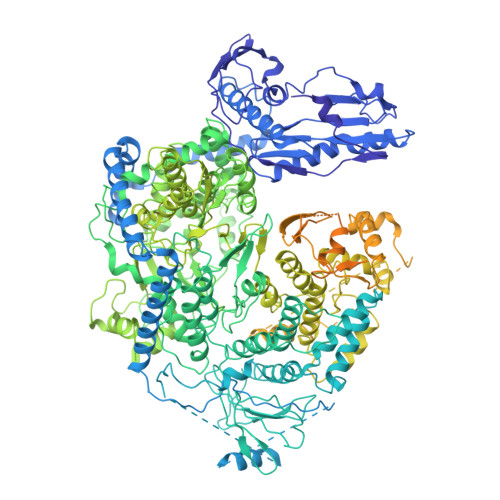
Structural and functional characterization of the severe fever with thrombocytopenia syndrome virus L protein.
Vogel, D., Thorkelsson, S.R., Quemin, E.R.J., Meier, K., Kouba, T., Gogrefe, N., Busch, C., Reindl, S., Gunther, S., Cusack, S., Grunewald, K., Rosenthal, M.(2020) Nucleic Acids Res 48: 5749-5765
- PubMed: 32313945
- DOI: https://doi.org/10.1093/nar/gkaa253
- Primary Citation of Related Structures:
6XYA, 6Y6K - PubMed Abstract:
The Bunyavirales order contains several emerging viruses with high epidemic potential, including Severe fever with thrombocytopenia syndrome virus (SFTSV). The lack of medical countermeasures, such as vaccines and antivirals, is a limiting factor for the containment of any virus outbreak. To develop such antivirals a profound understanding of the viral replication process is essential. The L protein of bunyaviruses is a multi-functional and multi-domain protein performing both virus transcription and genome replication and, therefore, is an ideal drug target. We established expression and purification procedures for the full-length L protein of SFTSV. By combining single-particle electron cryo-microscopy and X-ray crystallography, we obtained 3D models covering ∼70% of the SFTSV L protein in the apo-conformation including the polymerase core region, the endonuclease and the cap-binding domain. We compared this first L structure of the Phenuiviridae family to the structures of La Crosse peribunyavirus L protein and influenza orthomyxovirus polymerase. Together with a comprehensive biochemical characterization of the distinct functions of SFTSV L protein, this work provides a solid framework for future structural and functional studies of L protein-RNA interactions and the development of antiviral strategies against this group of emerging human pathogens.
Organizational Affiliation:
Department of Virology, Bernhard-Nocht-Institute for Tropical Medicine, Hamburg, Hamburg 20359, Germany.