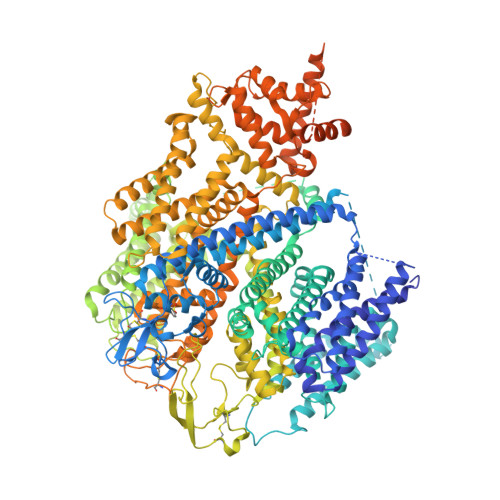
Structure of the human sodium leak channel NALCN.
Kschonsak, M., Chua, H.C., Noland, C.L., Weidling, C., Clairfeuille, T., Bahlke, O.O., Ameen, A.O., Li, Z.R., Arthur, C.P., Ciferri, C., Pless, S.A., Payandeh, J.(2020) Nature 587: 313-318
- PubMed: 32698188
- DOI: https://doi.org/10.1038/s41586-020-2570-8
- Primary Citation of Related Structures:
6XIW - PubMed Abstract:
Persistently depolarizing sodium (Na + ) leak currents enhance electrical excitability 1,2 . The ion channel responsible for the major background Na + conductance in neurons is the Na + leak channel, non-selective (NALCN) 3,4 . NALCN-mediated currents regulate neuronal excitability linked to respiration, locomotion and circadian rhythm 4-10 . NALCN activity is under tight regulation 11-14 and mutations in NALCN cause severe neurological disorders and early death 15,16 . NALCN is an orphan channel in humans, and fundamental aspects of channel assembly, gating, ion selectivity and pharmacology remain obscure. Here we investigate this essential leak channel and determined the structure of NALCN in complex with a distinct auxiliary subunit, family with sequence similarity 155 member A (FAM155A). FAM155A forms an extracellular dome that shields the ion-selectivity filter from neurotoxin attack. The pharmacology of NALCN is further delineated by a walled-off central cavity with occluded lateral pore fenestrations. Unusual voltage-sensor domains with asymmetric linkages to the pore suggest mechanisms by which NALCN activity is modulated. We found a tightly closed pore gate in NALCN where the majority of missense patient mutations cause gain-of-function phenotypes that cluster around the S6 gate and distinctive π-bulges. Our findings provide a framework to further study the physiology of NALCN and a foundation for discovery of treatments for NALCN channelopathies and other electrical disorders.
Organizational Affiliation:
Department of Structural Biology, Genentech Inc., San Francisco, CA, USA.