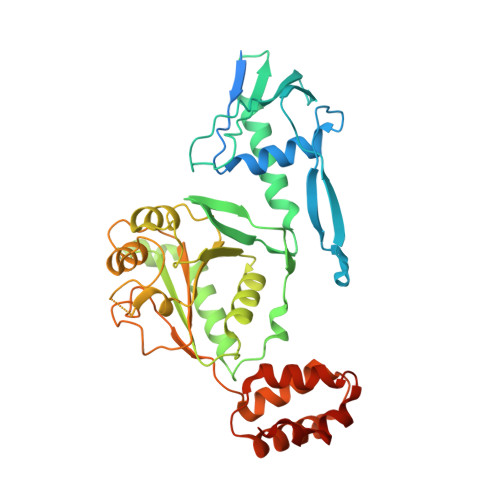
Structures of AAA protein translocase Bcs1 suggest translocation mechanism of a folded protein.
Tang, W.K., Borgnia, M.J., Hsu, A.L., Esser, L., Fox, T., de Val, N., Xia, D.(2020) Nat Struct Mol Biol 27: 202-209
- PubMed: 32042153
- DOI: https://doi.org/10.1038/s41594-020-0373-0
- Primary Citation of Related Structures:
6U1Y, 6UKO, 6UKP, 6UKS - PubMed Abstract:
The mitochondrial membrane-bound AAA protein Bcs1 translocate substrates across the mitochondrial inner membrane without previous unfolding. One substrate of Bcs1 is the iron-sulfur protein (ISP), a subunit of the respiratory Complex III. How Bcs1 translocates ISP across the membrane is unknown. Here we report structures of mouse Bcs1 in two different conformations, representing three nucleotide states. The apo and ADP-bound structures reveal a homo-heptamer and show a large putative substrate-binding cavity accessible to the matrix space. ATP binding drives a contraction of the cavity by concerted motion of the ATPase domains, which could push substrate across the membrane. Our findings shed light on the potential mechanism of translocating folded proteins across a membrane, offer insights into the assembly process of Complex III and allow mapping of human disease-associated mutations onto the Bcs1 structure.
Organizational Affiliation:
Laboratory of Cell Biology, National Cancer Institute, National Institutes of Health, Bethesda, MD, USA.