Molecular mechanism for the recognition of sequence-divergent CIF peptides by the plant receptor kinases GSO1/SGN3 and GSO2.
Okuda, S., Fujita, S., Moretti, A., Hohmann, U., Doblas, V.G., Ma, Y., Pfister, A., Brandt, B., Geldner, N., Hothorn, M.(2020) Proc Natl Acad Sci U S A 117: 2693-2703
- PubMed: 31964818
- DOI: https://doi.org/10.1073/pnas.1911553117
- Primary Citation of Related Structures:
6S6Q - PubMed Abstract:
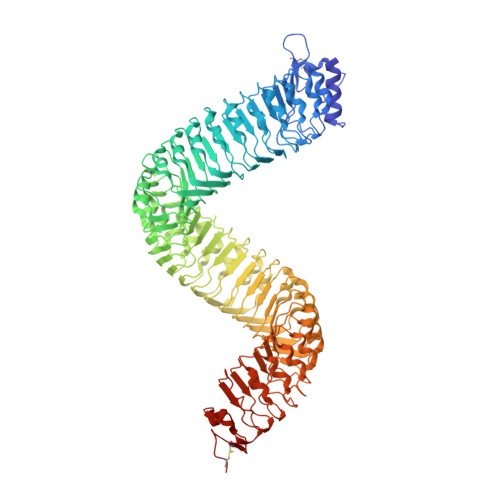
Plants use leucine-rich repeat receptor kinases (LRR-RKs) to sense sequence diverse peptide hormones at the cell surface. A 3.0-Å crystal structure of the LRR-RK GSO1/SGN3 regulating Casparian strip formation in the endodermis reveals a large spiral-shaped ectodomain. The domain provides a binding platform for 21 amino acid CIF peptide ligands, which are tyrosine sulfated by the tyrosylprotein sulfotransferase TPST/SGN2. GSO1/SGN3 harbors a binding pocket for sulfotyrosine and makes extended backbone interactions with CIF2. Quantitative biochemical comparisons reveal that GSO1/SGN3-CIF2 represents one of the strongest receptor-ligand pairs known in plants. Multiple missense mutations are required to block CIF2 binding in vitro and GSO1/SGN3 function in vivo. Using structure-guided sequence analysis we uncover previously uncharacterized CIF peptides conserved among higher plants. Quantitative binding assays with known and novel CIFs suggest that the homologous LRR-RKs GSO1/SGN3 and GSO2 have evolved unique peptide binding properties to control different developmental processes. A quantitative biochemical interaction screen, a CIF peptide antagonist and genetic analyses together implicate SERK proteins as essential coreceptor kinases required for GSO1/SGN3 and GSO2 receptor activation. Our work provides a mechanistic framework for the recognition of sequence-divergent peptide hormones in plants.
Organizational Affiliation:
Structural Plant Biology Laboratory, Department of Botany and Plant Biology, University of Geneva, 1211 Geneva, Switzerland.