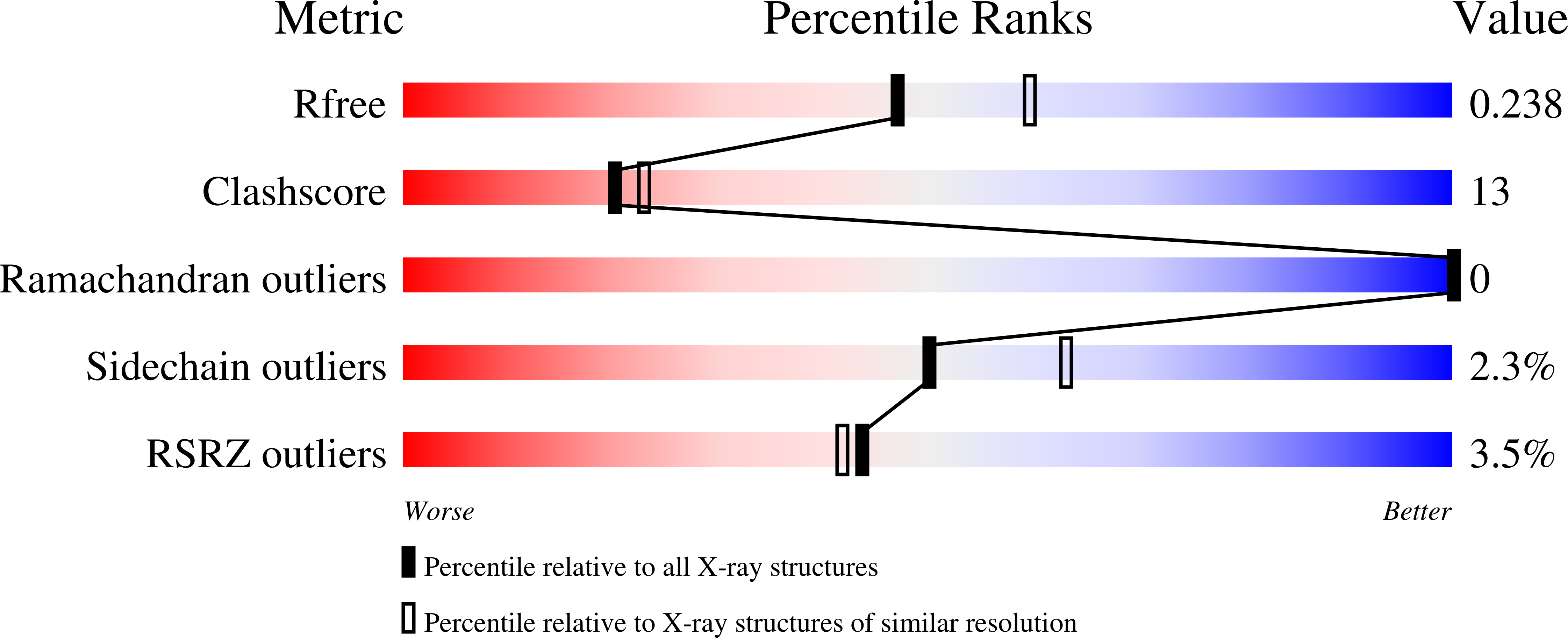
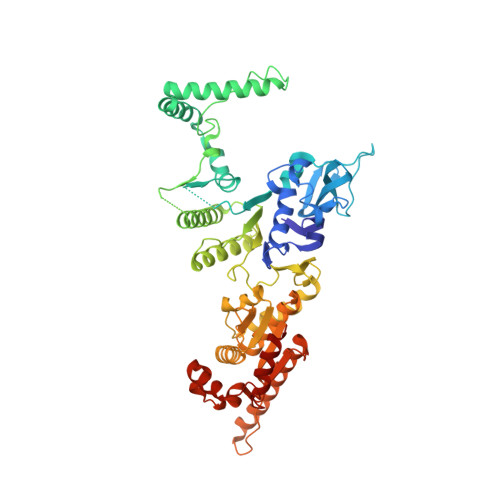
The full-length structure of Thermus scotoductus OLD defines the ATP hydrolysis properties and catalytic mechanism of Class 1 OLD family nucleases.
Schiltz, C.J., Adams, M.C., Chappie, J.S.(2020) Nucleic Acids Res 48: 2762-2776
- PubMed: 32009148
- DOI: https://doi.org/10.1093/nar/gkaa059
- Primary Citation of Related Structures:
6P74 - PubMed Abstract:
OLD family nucleases contain an N-terminal ATPase domain and a C-terminal Toprim domain. Homologs segregate into two classes based on primary sequence length and the presence/absence of a unique UvrD/PcrA/Rep-like helicase gene immediately downstream in the genome. Although we previously defined the catalytic machinery controlling Class 2 nuclease cleavage, degenerate conservation of the C-termini between classes precludes pinpointing the analogous residues in Class 1 enzymes by sequence alignment alone. Our Class 2 structures also provide no information on ATPase domain architecture and ATP hydrolysis. Here we present the full-length structure of the Class 1 OLD nuclease from Thermus scotoductus (Ts) at 2.20 Å resolution, which reveals a dimerization domain inserted into an N-terminal ABC ATPase fold and a C-terminal Toprim domain. Structural homology with genome maintenance proteins identifies conserved residues responsible for Ts OLD ATPase activity. Ts OLD lacks the C-terminal helical domain present in Class 2 OLD homologs yet preserves the spatial organization of the nuclease active site, arguing that OLD proteins use a conserved catalytic mechanism for DNA cleavage. We also demonstrate that mutants perturbing ATP hydrolysis or DNA cleavage in vitro impair P2 OLD-mediated killing of recBC-Escherichia coli hosts, indicating that both the ATPase and nuclease activities are required for OLD function in vivo.
Organizational Affiliation:
Department of Molecular Medicine, Cornell University, Ithaca, NY 14853, USA.