Structures of teixobactin-producing nonribosomal peptide synthetase condensation and adenylation domains.
Tan, K., Zhou, M., Jedrzejczak, R.P., Wu, R., Higuera, R.A., Borek, D., Babnigg, G., Joachimiak, A.(2020) Curr Res Struct Biol 2: 14-24
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
(2020) Curr Res Struct Biol 2: 14-24
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Txo2 | 964 | Eleftheria terrae | Mutation(s): 0 | 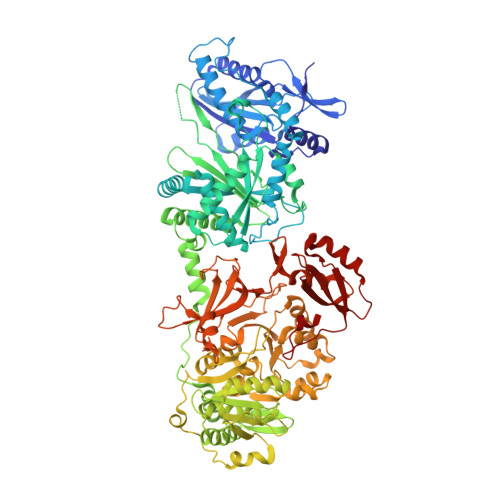 | |
UniProt | |||||
Find proteins for A0A0B5H0S3 (Eleftheria terrae) Explore A0A0B5H0S3 Go to UniProtKB: A0A0B5H0S3 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A0B5H0S3 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| FLC Query on FLC | D [auth A], W [auth B] | CITRATE ANION C6 H5 O7 KRKNYBCHXYNGOX-UHFFFAOYSA-K |  | ||
| SO4 Query on SO4 | Q [auth A], R [auth A] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| GOL Query on GOL | L [auth A], M [auth A], N [auth A], O [auth A], P [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| ACT Query on ACT | E [auth A] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| FMT Query on FMT | C [auth A], S [auth B], T [auth B], U [auth B], V [auth B] | FORMIC ACID C H2 O2 BDAGIHXWWSANSR-UHFFFAOYSA-N |  | ||
| CL Query on CL | AA [auth B] BA [auth B] CA [auth B] DA [auth B] EA [auth B] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 110.857 | α = 90 |
| b = 399.875 | β = 90 |
| c = 144.116 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-3000 | data reduction |
| HKL-3000 | data scaling |
| HKL-3000 | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID) | United States | HHSN272201700060C |