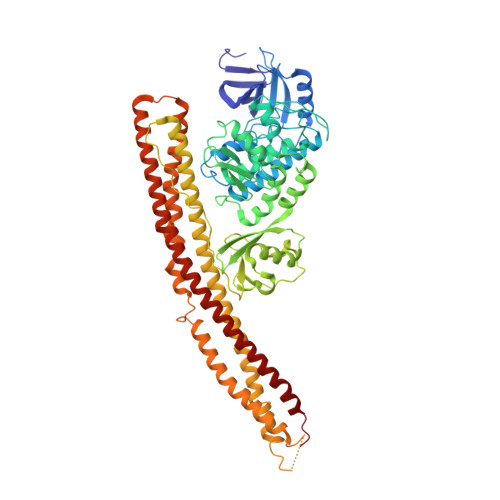
A conserved PLPLRT/SD motif of STING mediates the recruitment and activation of TBK1.
Zhao, B., Du, F., Xu, P., Shu, C., Sankaran, B., Bell, S.L., Liu, M., Lei, Y., Gao, X., Fu, X., Zhu, F., Liu, Y., Laganowsky, A., Zheng, X., Ji, J.Y., West, A.P., Watson, R.O., Li, P.(2019) Nature 569: 718-722
- PubMed: 31118511
- DOI: https://doi.org/10.1038/s41586-019-1228-x
- Primary Citation of Related Structures:
6O8B, 6O8C - PubMed Abstract:
Nucleic acids from bacteria or viruses induce potent immune responses in infected cells 1-4 . The detection of pathogen-derived nucleic acids is a central strategy by which the host senses infection and initiates protective immune responses 5,6 . Cyclic GMP-AMP synthase (cGAS) is a double-stranded DNA sensor 7,8 . It catalyses the synthesis of cyclic GMP-AMP (cGAMP) 9-12 , which stimulates the induction of type I interferons through the STING-TBK1-IRF-3 signalling axis 13-15 . STING oligomerizes after binding of cGAMP, leading to the recruitment and activation of the TBK1 kinase 8,16 . The IRF-3 transcription factor is then recruited to the signalling complex and activated by TBK1 8,17-20 . Phosphorylated IRF-3 translocates to the nucleus and initiates the expression of type I interferons 21 . However, the precise mechanisms that govern activation of STING by cGAMP and subsequent activation of TBK1 by STING remain unclear. Here we show that a conserved PLPLRT/SD motif within the C-terminal tail of STING mediates the recruitment and activation of TBK1. Crystal structures of TBK1 bound to STING reveal that the PLPLRT/SD motif binds to the dimer interface of TBK1. Cell-based studies confirm that the direct interaction between TBK1 and STING is essential for induction of IFNβ after cGAMP stimulation. Moreover, we show that full-length STING oligomerizes after it binds cGAMP, and highlight this as an essential step in the activation of STING-mediated signalling. These findings provide a structural basis for the development of STING agonists and antagonists for the treatment of cancer and autoimmune disorders.
Organizational Affiliation:
Department of Biochemistry and Biophysics, Texas A&M University, College Station, TX, USA.