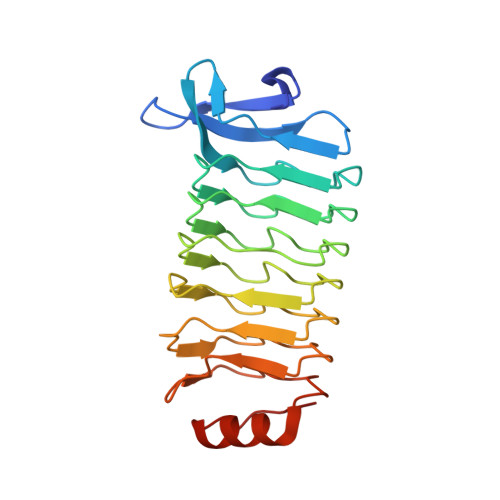
Structural characterization of Treponema pallidum Tp0225 reveals an unexpected leucine-rich repeat architecture.
Ramaswamy, R., Houston, S., Loveless, B., Cameron, C.E., Boulanger, M.J.(2019) Acta Crystallogr F Struct Biol Commun 75: 489-495
- PubMed: 31282868
- DOI: https://doi.org/10.1107/S2053230X19007726
- Primary Citation of Related Structures:
6MLX - PubMed Abstract:
The phylogenetically divergent spirochete bacterium Treponema pallidum subsp. pallidum is the causative agent of syphilis. Central to the capacity of T. pallidum to establish infection is the ability of the pathogen to attach to a diversity of host cells. Many pathogenic bacteria employ leucine-rich repeat (LRR) domain-containing proteins to mediate protein-protein interactions, including attachment to host components and establishment of infection. Intriguingly, T. pallidum expresses only one putative LRR domain-containing protein (Tp0225) with an unknown function. In an effort to ascribe a function to Tp0225, a comprehensive phylogenetic analysis was first performed; this investigation revealed that Tp0225 clusters with the pathogenic clade of treponemes. Its crystal structure was then determined to 2.0 Å resolution using Pt SAD phasing, which revealed a noncanonical architecture containing a hexameric LRR core with a discontinuous β-sheet bridged by solvent molecules. Furthermore, a surface-exposed, hydrophobic pocket, which was found in Tp0225 but is largely absent in canonical LRR domains from other pathogenic bacteria, may serve to coordinate a hydrophobic ligand. Overall, this study provides the first structural characterization of the sole LRR domain-containing protein from T. pallidum and offers insight into the unique molecular landscape of this important human pathogen.
Organizational Affiliation:
Department of Biochemistry and Microbiology, University of Victoria, Victoria, BC V8P 5C2, Canada.