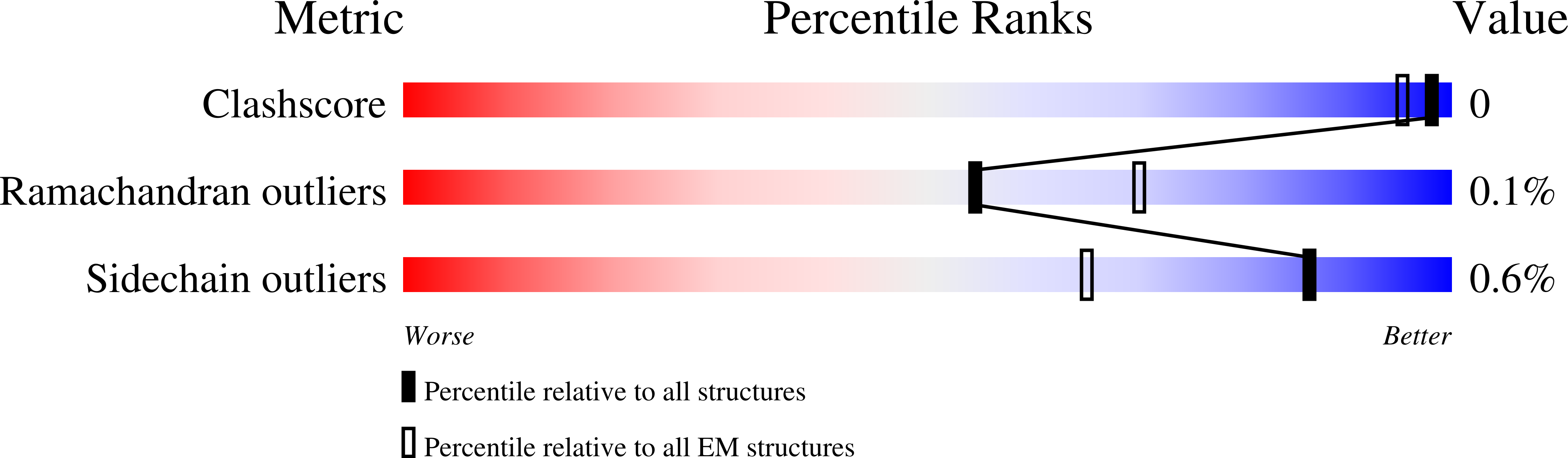Cryo-EM structure ofP. falciparumcircumsporozoite protein with a vaccine-elicited antibody is stabilized by somatically mutated inter-Fab contacts.
Oyen, D., Torres, J.L., Cottrell, C.A., Richter King, C., Wilson, I.A., Ward, A.B.(2018) Sci Adv 4: eaau8529-eaau8529
- PubMed: 30324137
- DOI: https://doi.org/10.1126/sciadv.aau8529
- Primary Citation of Related Structures:
6MB3, 6MHG - PubMed Abstract:
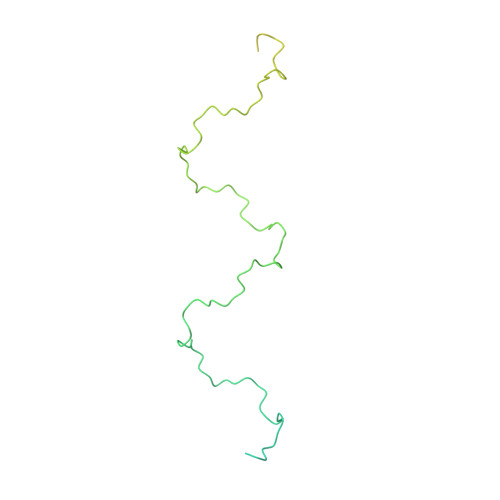
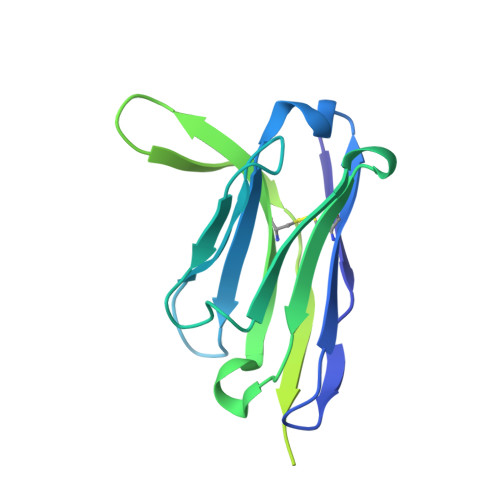
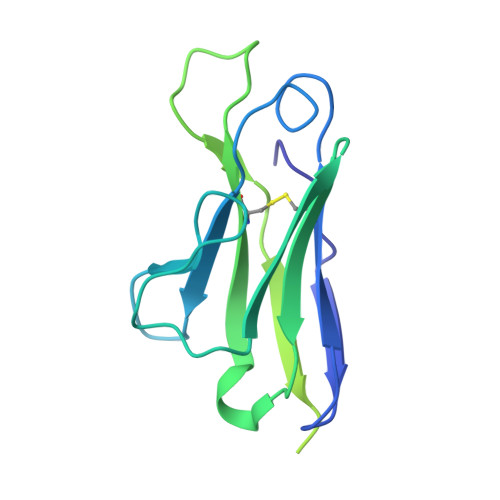
The circumsporozoite protein (CSP) on the surface of Plasmodium falciparum sporozoites is important for parasite development, motility, and host hepatocyte invasion. However, intrinsic disorder of the NANP repeat sequence in the central region of CSP has hindered its structural and functional characterization. Here, the cryo-electron microscopy structure at ~3.4-Å resolution of a recombinant shortened CSP construct with the variable domains (Fabs) of a highly protective monoclonal antibody reveals an extended spiral conformation of the central NANP repeat region surrounded by antibodies. This unusual structure appears to be stabilized and/or induced by interaction with an antibody where contacts between adjacent Fabs are somatically mutated and enhance the interaction. This maturation in non-antigen contact residues may be an effective mechanism for antibodies to target tandem repeat sequences and provide novel insights into malaria vaccine design.
Organizational Affiliation:
Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.