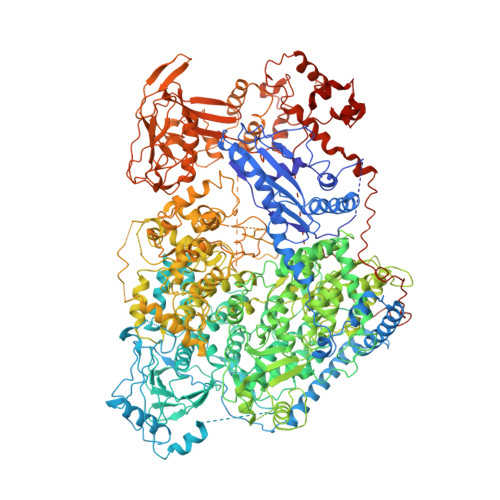
Structure of severe fever with thrombocytopenia syndrome virus L protein elucidates the mechanisms of viral transcription initiation.
Wang, P., Liu, L., Liu, A., Yan, L., He, Y., Shen, S., Hu, M., Guo, Y., Liu, H., Liu, C., Lu, Y., Wang, P., Deng, F., Rao, Z., Lou, Z.(2020) Nat Microbiol 5: 864-871
- PubMed: 32341479
- DOI: https://doi.org/10.1038/s41564-020-0712-2
- Primary Citation of Related Structures:
6L42 - PubMed Abstract:
Segmented negative-sense RNA viruses (sNSRVs) encode a single-polypeptide polymerase (L protein) or a heterotrimeric polymerase complex to cannibalize host messenger RNA cap structures serving as primers of transcription, and catalyse RNA synthesis. Here, we report the full-length structure of the severe fever with thrombocytopaenia syndrome virus (SFTSV) L protein, as determined by cryogenic electron microscopy at 3.4 Å, leading to an atomic model harbouring three functional parts (an endonuclease, an RNA-dependent RNA polymerase and a cap-binding domain) and two structural domains (an arm domain with a blocker motif and a carboxy-terminal lariat domain). The SFTSV L protein has a compact architecture in which its cap-binding pocket is surprisingly occupied by an Arg finger of the blocker motif, and the endonuclease active centre faces back towards the cap-binding pocket, suggesting that domain rearrangements are necessary to acquire the pre-initiation state of the active site. Our results provide insight into the complete architecture of sNSRV-encoded L protein and further the understanding of sNSRV transcription initiation.
Organizational Affiliation:
MOE Key Laboratory of Protein Science and Collaborative Innovation Center of Biotherapy, School of Medicine, Tsinghua University, Beijing, China.