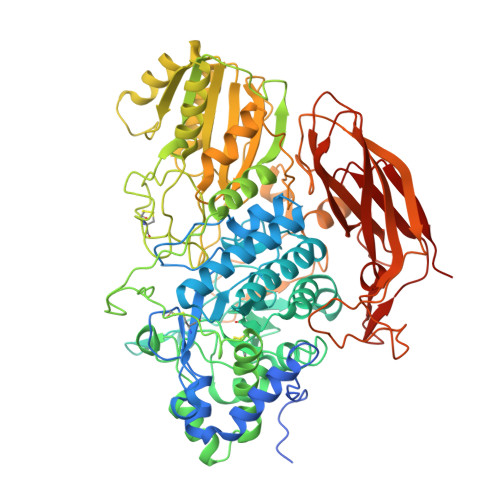
Structures of beta-glycosidase LXYL-P1-2 reveals the product binding state of GH3 family and a specific pocket for Taxol recognition.
Yang, L.Y., Chen, T.J., Wang, F., Li, L., Yu, W.B., Si, Y.K., Chen, J.J., Liu, W.C., Zhu, P., Gong, W.M.(2020) Commun Biol 3: 22-22
- PubMed: 31925310
- DOI: https://doi.org/10.1038/s42003-019-0744-4
- Primary Citation of Related Structures:
6JBS, 6KJ0 - PubMed Abstract:
LXYL-P1-2 is one of the few xylosidases that efficiently catalyze the reaction from 7-β-xylosyl-10-deacetyltaxol (XDT) to 10-deacetyltaxol (DT), and is a potential enzyme used in Taxol industrial production. Here we report the crystal structure of LXYL-P1-2 and its XDT binding complex. These structures reveal an enzyme/product complex with the sugar conformation different from the enzyme/substrate complex reported previously in GH3 enzymes, even in the whole glycohydrolases family. In addition, the DT binding pocket is identified as the structural basis for the substrate specificity. Further structure analysis reveals common features in LXYL-P1-2 and Taxol binding protein tubulin, which might provide useful information for designing new Taxol carrier proteins for drug delivery.
Organizational Affiliation:
National Research Laboratory for Physical Sciences in Microscales, University of Science and Technology of China, 230026, Hefei, Anhui, China.