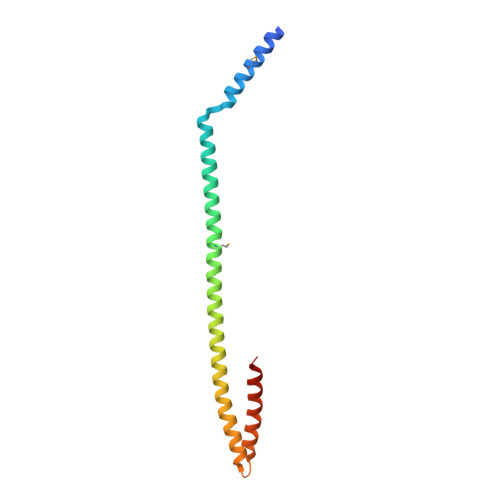
Crystal structure of the coiled-coil domain of Drosophila TRIM protein Brat.
Liu, C., Shan, Z., Diao, J., Wen, W., Wang, W.(2019) Proteins 87: 706-710
- PubMed: 30958583
- DOI: https://doi.org/10.1002/prot.25691
- Primary Citation of Related Structures:
6J9R - PubMed Abstract:
Drosophila brain tumor (Brat) is a translational repressor belonging to the tripartite motif (TRIM) protein superfamily. During the asymmetric division of Drosophila neuroblasts, Brat localizes at the basal cortex via direct interaction with the scaffolding protein Miranda (Mira), and segregates into the basal ganglion mother cells after cell division. It was previously reported that both the coiled-coil (CC) and NHL domains of Brat are required for the interaction with Mira, but the underlying structural basis is elusive. Here, we determine the crystal structure of Brat-CC domain (aa 376-511) at 2.5 Å, showing that Brat-CC forms an elongated antiparallel dimer through an unconventional CC structure. The dimeric assembly in Brat-CC structure is similar to its counterparts in other TRIM proteins, but Brat-CC also exhibits some distinct structural features. We also demonstrate that the CC domain could not bind Mira by its own, neither does the isolated NHL domain of Brat. Rather, Brat binds to Mira through the CC-NHL domain tandem, indicating that the function of the CC domain is to assemble Brat-NHL in dimeric form, which is necessary for Mira binding.
Organizational Affiliation:
Department of Chemistry, Institutes of Biomedical Sciences and Multiscale Research Institute of Complex System, Fudan University, Shanghai, People's Republic of China.