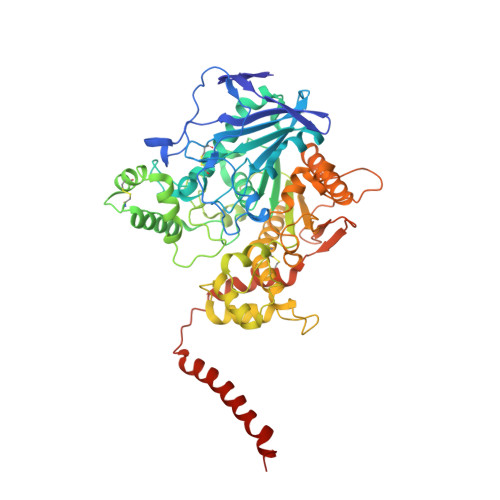
Cryo-EM structure of the native butyrylcholinesterase tetramer reveals a dimer of dimers stabilized by a superhelical assembly.
Leung, M.R., van Bezouwen, L.S., Schopfer, L.M., Sussman, J.L., Silman, I., Lockridge, O., Zeev-Ben-Mordehai, T.(2018) Proc Natl Acad Sci U S A 115: 13270-13275
- PubMed: 30538207
- DOI: https://doi.org/10.1073/pnas.1817009115
- Primary Citation of Related Structures:
6I2T - PubMed Abstract:
The quaternary structures of the cholinesterases, acetylcholinesterase (AChE) and butyrylcholinesterase (BChE), are essential for their localization and function. Of practical importance, BChE is a promising therapeutic candidate for intoxication by organophosphate nerve agents and insecticides, and for detoxification of addictive substances. Efficacy of the recombinant enzyme hinges on its having a long circulatory half-life; this, in turn, depends strongly on its ability to tetramerize. Here, we used cryoelectron microscopy (cryo-EM) to determine the structure of the highly glycosylated native BChE tetramer purified from human plasma at 5.7 Å. Our structure reveals that the BChE tetramer is organized as a staggered dimer of dimers. Tetramerization is mediated by assembly of the C-terminal tryptophan amphiphilic tetramerization (WAT) helices from each subunit as a superhelical assembly around a central lamellipodin-derived oligopeptide with a proline-rich attachment domain (PRAD) sequence that adopts a polyproline II helical conformation and runs antiparallel. The catalytic domains within a dimer are asymmetrically linked to the WAT/PRAD. In the resulting arrangement, the tetramerization domain is largely shielded by the catalytic domains, which may contribute to the stability of the human BChE (HuBChE) tetramer. Our cryo-EM structure reveals the basis for assembly of the native tetramers and has implications for the therapeutic applications of HuBChE. This mode of tetramerization is seen only in the cholinesterases but may provide a promising template for designing other proteins with improved circulatory residence times.
Organizational Affiliation:
Cryo-Electron Microscopy, Bijvoet Center for Biomolecular Research, Utrecht University, 3584 CH Utrecht, The Netherlands.