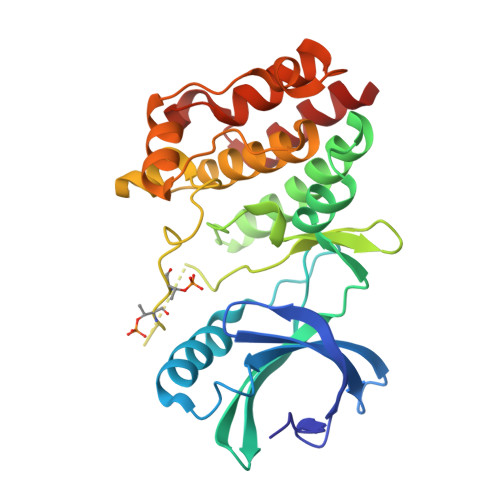
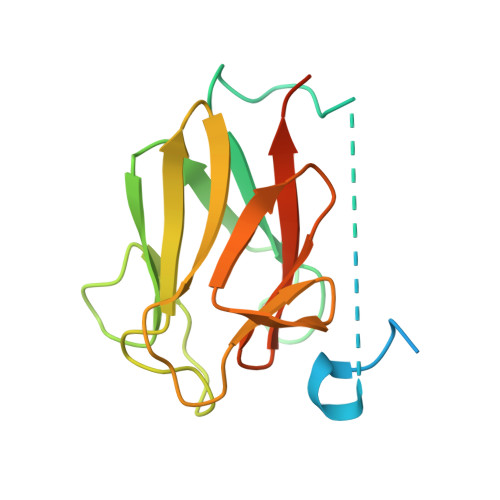
Structural insights into the functional versatility of an FHA domain protein in mycobacterial signaling.
Wagner, T., Andre-Leroux, G., Hindie, V., Barilone, N., Lisa, M.N., Hoos, S., Raynal, B., Vulliez-Le Normand, B., O'Hare, H.M., Bellinzoni, M., Alzari, P.M.(2019) Sci Signal 12
- PubMed: 31064884
- DOI: https://doi.org/10.1126/scisignal.aav9504
- Primary Citation of Related Structures:
6I2P, 6I2Q, 6I2R, 6I2S - PubMed Abstract:
Forkhead-associated (FHA) domains are modules that bind to phosphothreonine (pThr) residues in signaling cascades. The FHA-containing mycobacterial protein GarA is a central element of a phosphorylation-dependent signaling pathway that redirects metabolic flux in response to amino acid starvation or cell growth requirements. GarA acts as a phosphorylation-dependent ON/OFF molecular switch. In its nonphosphorylated ON state, the GarA FHA domain engages in phosphorylation-independent interactions with various metabolic enzymes that orchestrate nitrogen flow, such as 2-oxoglutarate decarboxylase (KGD). However, phosphorylation at the GarA N-terminal region by the protein kinase PknB or PknG triggers autoinhibition through the intramolecular association of the N-terminal domain with the FHA domain, thus blocking all downstream interactions. To investigate these different FHA binding modes, we solved the crystal structures of the mycobacterial upstream (phosphorylation-dependent) complex PknB-GarA and the downstream (phosphorylation-independent) complex GarA-KGD. Our results show that the phosphorylated activation loop of PknB serves as a docking site to recruit GarA through canonical FHA-pThr interactions. However, the same GarA FHA-binding pocket targets an allosteric site on nonphosphorylated KGD, where a key element of recognition is a phosphomimetic aspartate. Further enzymatic and mutagenesis studies revealed that GarA acted as a dynamic allosteric inhibitor of KGD by preventing crucial motions in KGD that are necessary for catalysis. Our results provide evidence for physiological phosphomimetics, supporting numerous mutagenesis studies using such approaches, and illustrate how evolution can shape a single FHA-binding pocket to specifically interact with multiple phosphorylated and nonphosphorylated protein partners.
Organizational Affiliation:
Institut Pasteur, Unité de Microbiologie Structurale, CNRS UMR 3528 & Université Paris Diderot, 25 Rue du Docteur Roux, 75724 Paris Cedex 15, France.