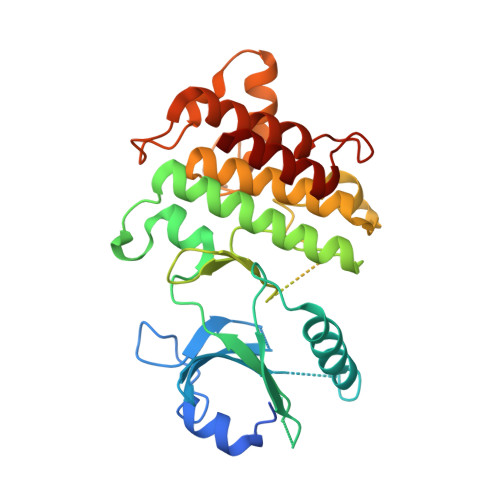
Conformational flexibility and inhibitor binding to unphosphorylated interleukin-1 receptor-associated kinase 4 (IRAK4).
Wang, L., Ferrao, R., Li, Q., Hatcher, J.M., Choi, H.G., Buhrlage, S.J., Gray, N.S., Wu, H.(2019) J Biol Chem 294: 4511-4519
- PubMed: 30679311
- DOI: https://doi.org/10.1074/jbc.RA118.005428
- Primary Citation of Related Structures:
6EG9, 6EGA, 6EGD, 6EGE, 6EGF - PubMed Abstract:
Interleukin-1 receptor-associated kinase 4 (IRAK4) is a key player in innate immune and inflammatory responses, performing a critical role in signal transduction downstream of Toll-like receptors and interleukin-1 (IL-1) receptors. Upon ligand binding and via its N-terminal death domain, IRAK4 is recruited to an oligomeric receptor that is proximal to the Myddosome signaling complex, inducing IRAK4 kinase domain dimerization, autophosphorylation, and activation. To date, all known IRAK4 structures are in the active conformation, precluding a good understanding of IRAK4's conformational dynamics. To address this issue, here we first solved three crystal structures of the IRAK4 kinase domain (at ≤2.6 Å resolution), in its unphosphorylated, inactive state bound to either the ATP analog AMP-PNP or to one of the two small-molecule inhibitors JH-I-25 and JH-I-17. The structures disclosed that although the structure in complex with AMP-PNP is in an "αC-out" inactive conformation, those in complex with type I inhibitors assume an active "Asp-Phe-Gly (DFG)-in" and "αC-in" conformation. The ability of unphosphorylated IRAK4 to take on variable conformations prompted us to screen for small-molecule inhibitors that bind preferentially to unphosphorylated IRAK4, leading to the identification of ponatinib and HG-12-6. Solving the structures of unphosphorylated IRAK4 in complex with these two inhibitors, we found that they both bind as type II inhibitors with IRAK4 in a "DFG-out" conformation. Collectively, these structures reveal conformational flexibility of unphosphorylated IRAK4 and provide unexpected insights into the potential use of small molecules to modulate IRAK4 activity in cancer, autoimmunity, and inflammation.
Organizational Affiliation:
From the Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, Massachusetts 02115.