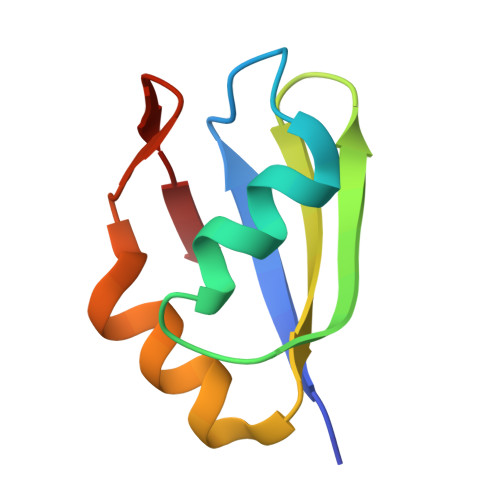
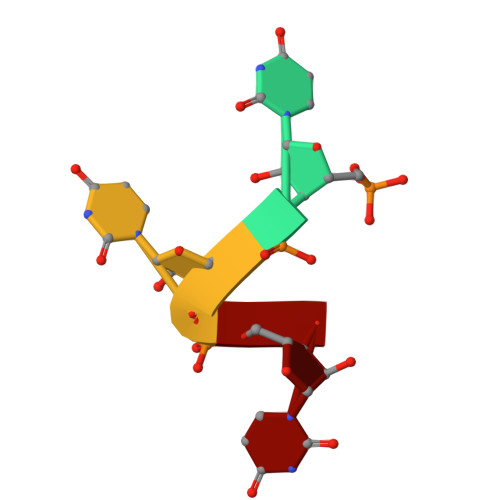
The RRM of the kRNA-editing protein TbRGG2 uses multiple surfaces to bind and remodel RNA.
Travis, B., Shaw, P.L.R., Liu, B., Ravindra, K., Iliff, H., Al-Hashimi, H.M., Schumacher, M.A.(2019) Nucleic Acids Res 47: 2130-2142
- PubMed: 30544166
- DOI: https://doi.org/10.1093/nar/gky1259
- Primary Citation of Related Structures:
6E4N, 6E4O, 6E4P - PubMed Abstract:
Kinetoplastid RNA (kRNA) editing takes place in the mitochondria of kinetoplastid protists and creates translatable mRNAs by uridine insertion/deletion. Extensively edited (pan-edited) transcripts contain quadruplex forming guanine stretches, which must be remodeled to promote uridine insertion/deletion. Here we show that the RRM domain of the essential kRNA-editing factor TbRGG2 binds poly(G) and poly(U) RNA and can unfold both. A region C-terminal to the RRM mediates TbRGG2 dimerization, enhancing RNA binding. A RRM-U4 RNA structure reveals a unique RNA-binding mechanism in which the two RRMs of the dimer employ aromatic residues outside the canonical RRM RNA-binding motifs to encase and wrench open the RNA, while backbone atoms specify the uridine bases. Notably, poly(G) RNA is bound via a different binding surface. Thus, these data indicate that TbRGG2 RRM can bind and remodel several RNA substrates suggesting how it might play multiple roles in the kRNA editing process.
Organizational Affiliation:
Department of Biochemistry, Duke University School of Medicine, Durham, NC 27710, USA.