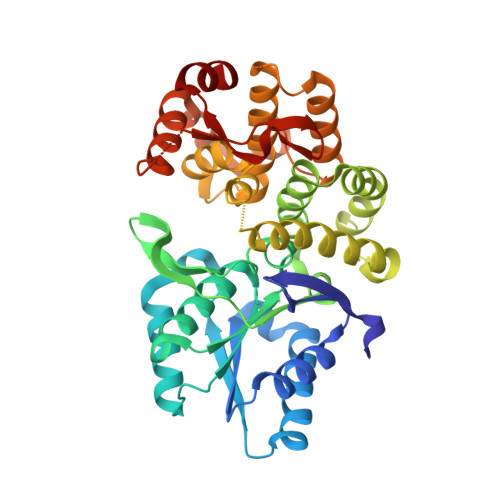
Molecular analysis and essentiality of Aro1 shikimate biosynthesis multi-enzyme in Candida albicans.
Stogios, P.J., Liston, S.D., Semper, C., Quade, B., Michalska, K., Evdokimova, E., Ram, S., Otwinowski, Z., Borek, D., Cowen, L.E., Savchenko, A.(2022) Life Sci Alliance 5
- PubMed: 35512834
- DOI: https://doi.org/10.26508/lsa.202101358
- Primary Citation of Related Structures:
6C5C, 7TBU, 7TBV, 7U5S, 7U5T, 7U5U - PubMed Abstract:
In the human fungal pathogen Candida albicans , ARO1 encodes an essential multi-enzyme that catalyses consecutive steps in the shikimate pathway for biosynthesis of chorismate, a precursor to folate and the aromatic amino acids. We obtained the first molecular image of C. albicans Aro1 that reveals the architecture of all five enzymatic domains and their arrangement in the context of the full-length protein. Aro1 forms a flexible dimer allowing relative autonomy of enzymatic function of the individual domains. Our activity and in cellulo data suggest that only four of Aro1's enzymatic domains are functional and essential for viability of C. albicans , whereas the 3-dehydroquinate dehydratase (DHQase) domain is inactive because of active site substitutions. We further demonstrate that in C. albicans , the type II DHQase Dqd1 can compensate for the inactive DHQase domain of Aro1, suggesting an unrecognized essential role for this enzyme in shikimate biosynthesis. In contrast, in Candida glabrata and Candida parapsilosis , which do not encode a Dqd1 homolog, Aro1 DHQase domains are enzymatically active, highlighting diversity across Candida species.
Organizational Affiliation:
Department of Chemical Engineering and Applied Chemistry, University of Toronto, Toronto, Canada.